Kamagra gibt es auch als Kautabletten, die sich schneller auflösen als normale Pillen. Manche Patienten empfinden das als angenehmer. Wer sich informieren will, findet Hinweise unter kamagra kautabletten.
Chiriottieditori.it
Antibiotic susceptibility of potentiAlly
probiotic LactobaciLLus strAins
JunhuA hAna, DAhuAn chena, shAnshAn lic, Xingfeng lia,
Wen-Wen Zhoud, bolin ZhAngb,*, yingmin JiAa,*
aCollege of Biological Science and Engineering, Hebei University of Science and Technology,
Shijiazhuang, Heibei, 050018, China
bSchool of Biological Science and Biotechnology, Beijing Forestry University, Beijing, 100083, China
c Dongcheng District Center For Disease Control And Prevention, Beijing, 100009, China
dSchool of Biosystems Engineering and Food Science, Zhejiang University, Hangzhou Zhejiang,
*Corresponding authors:
Yingmin Jia, Tel. +86 311 81668010,
[email protected]
Bolin Zhang, Tel. +86 10 62338221,
susceptibility of 29
Lactobacilli to 13 antibiotics was assayed by paper disc diffusion method.
Plasmids and gastrointestinal tolerance were detected. the relationship between plasmids and
antibiotic resistance was discussed. the results showed that all of the strains were resistant to
bacitracin, polymyxin b, kanamycin, and nalidixic acid. Many strains were relatively sensitive to
chloramphenicol and tetracycline. six strains
contained plasmids and showed good gastrointesti-
nal tolerance. β-lactam resistance gene
blr was found in the plasmid of
L. plantarum cIcc 23180
by Pcr. the study will be helpful to promote the safety evaluation and development of potential-
ly probiotic lactic acid bacteria.
- Keywords: antibiotic resistance;
Lactobacillus; gastrointestinal tolerance; plasmid; probiotic -
282 Ital. J. Food Sci., vol. 27 - 2015
testinal conditions was investigated. by plas-
mid elimination and Pcr, the relationship be-
tween the plasmid profiles and resistance pat-
due to the claimed benefits,
Lactobacillus
terns of the strains was explored. this will pro-
bacteria are widely used in food, feed, medical
vide a reference for the safety evaluation meth-
and health related fields. Many lactic acid bac-
od and also will be helpful to improve the eval-
teria (LAb), such as
Streptococcus thermophi-
uation system of probiotics.
lus and
Lactobacillus delbruekii subsp.
bulga-
ricus, have been used safely for a long history.
they are agreed to be secure and do not have
2. MAtErIALs And MEtHods
the possibility of pathogenic. currently, new
beneficial bacteria are being developed contin-
2.1 Bacterial strains and cultivation
uously and will enter the market. However, the
security of these new strains has caused great
29
Lactobacillus strains used in the study
concern. Evaluation of antibiotic sensitivity is
were listed in table 1.
Lactobacillus strains
an important part of safety assessment.
were cultured in Mrs (de Man, rogosa, and
now, overuse of antibiotics has become a
sharpe) medium at 37°c for 18h under anaer-
serious social problem. this led to the emer-
obic condition.
gence of a large number of antibiotic-resistant
Quality control strain recommended by clin-
strains. once the resistance-related factors are
ical and Laboratory standards Institute (cLsI)
tranferred to other microorganisms, especial-
in the antibiotic sensitivity test was
E. coli
ly pathogens via food carrier, it will cause tre-
Atcc25922 purchased from the Institute of Mi-
mendous problems. the evolution of antibiotic-
crobiology, chinese Academy of sciences. the
resistant foodborne pathogens has been widely
E. coli Atcc25922 was activated and cultivat-
reported (tHrELfALL
et al., 2000; WALsH
et al.,
ed in Lb medium (yeast extract 5 g/L, tryptone
2008; WHItE
et al., 2002). Moreover, the resist-
10 g/L, nacl 10 g/L) at 37°c.
ance and resistance-related genes of
Bifidobac-
terium,
Lactobacillus and
Pediococcus strains to
2.2 Testing of antibiotic susceptibility
different antibiotics were studied systematical-
ly (HuMMEL
et al., 2007; Huys
et al., 2004; MA-
13 kinds of antibiotics paper discs were pur-
rIA
et al., 2007). the
tetM gene transfer of tet-
chased from the national Institute for the con-
racycline resistance in
Lactobacillus plantar-
trol of Pharmaceutical and biological Products
um among strains was reported by nIAMH
et
(table 2), each piece with a diameter of 6.5 mm.
al. (2010).
the quality was fully complied with the WHo
In this study, 29
Lactobacillus strains isolat-
ed from the food environment with potentially
Antibiotic susceptibility was semi-quantita-
probiotic effects (JIn
et al., 2009; LI
et al., 2009;
tively determined with K-b method by antibiot-
LIu
et al., 2011; sun
et al., 2009; ZHAo
et al.,
ic paper disc diffusion referring to the cLsI as
2013) were used. these strains were assayed for
described by cHArtErIs
et al. (1998a).
susceptibility to 13 antibiotics by agar disc dif-
briefly, 1.0 mL
Lactobacillus suspension (ap-
fusion method. furthermore, some strains with
proximately 1.5×108 cfu/mL) was added to
higher resistance were analysed for the presence
sterile petri dish with diameter of 90 mm, and
of plasmids. then, the tolerance of the plasmid-
then mixed with a 15 mL MH (Muller Hinton,
containing strains under simulated gastroin-
MH) agar (beef extract powder 6g/L, casein ac-
table 1 - source of the tested strains for antibiotic susceptibility test.
Source (original number)
Lactobacillus plantarum
CICCa 23124 (L11), CICC 23131 (B31), CICC 23135 (B37), CICC 22195 (C35),
CICC 23166 (ZJ1), CICC 23138 (C8-1), CICC 23180 (CH8)
Lactobacillus rhamnosus
CICC 23119 (1132), CICC 22175 (LL), ATCCb 7469, CICC 22151 (LK-Mt), CICC 22173 (R11)
Lactobacillus salivarius
CICC 23182 (CH-10)
Lactobacillus acidophilus
CICC 22162 (CH-2)
Lactobacillus casei
CICC 23184 (Y5-2b)
Lactobacillus helveticus
Lactobacillus pentosus
CICC 23116 (SN23), CICC 22161 (Lp-4), CICC 22160 (Lp-5), CICC 22159 (Lp-B),
CICC 22156 (Ind-3), CICC 22157 (Lp-A)
Lactobacillus paralimentarius
CICC 22148 (412), CICC 22149 (413)
Lactobacillus delbrueckii
CICC 22153 (LB), CICC 22163 (LC)
Lactobacillus paracasei
CICC 22165 (5M1), CICC 22167 (5M7), CICC 23183 (D-400)
aCICC, China center of industrial culture collection. bATCC, American type culture collection.
Ital. J. Food Sci., vol. 27 - 2015 283
table 2 - the content of antibiotic paper discs and criteri-
2.4 Gastrointestinal tolerability test
on for judgement.
Antibiotics
Content/disc inhibition zone diameter (mm)
In order to explore the application safety
in human, the gastrointestinal tolerability of
those lactic acid bacteria containing the plas-
mids were tested.
for acid tolerance test, Lactobacillus cells
≤14 15-17 ≥18
were harvested by centrifugation at 6000 rpm
μg ≤21 22-28 ≥29
for 15 min, washed twice with 0.01 mol/L Pbs,
pH 7.2 after cultured for 18 h at 37°c in Mrs
μg ≤14 15-17 ≥18
broth, and then suspended in 20 mL sterile sa-
line (0.85%, w/v) adjusted to pH 2.5 with ster-
μg ≤13 14-17 ≥18
ile hydrochloric acid.
for bile tolerance test, the modified method
chloramphenicol 30
μg ≤12 13-17 ≥18
of LEE
et al. (1999) was referred to test bile tol-
μg ≤12 13-14 ≥15
erance. the
Lactobacillus cells were centrifuged
(6000 rpm, 15 min) after cultivated for 18 h at
multi-polymyxin B
37°c in Mrs broth and suspended in 20 mL ster-
ile saline (0.85 %, w/v) supplemented with 0.3%
(w/v) bile salts (taurocholate, sigma) at pH 6.8.
cNote: R-Resistant; S-Susceptible; I-Intermediate.
for
pepsin and trypsin tolerance test,
Lactoba-
cillus cells were centrifuged (6000 rpm, 15 min)
after cultivated for 18 h at 37°c in Mrs broth,
then suspended in 20 mL sterile simulated gas-
ids hydrolysate 17.5 g/L, soluble starch, 1.5
tric and pancretic juices. fresh simulated gas-
g/L, agar 17 g/L, pH 7.3±0.1) until the medi-
tric and pancreatic juices were prepared daily
um solidified. the antibiotic paper discs were
according to charteris
et al (1998b). Pepsin (sig-
pasted closely onto the solidified medium with
ma) was added into the simulated gastric juice
sterile tweezers after 5min at room temper-
with a final concentration of 5 mg/mL. then the
ature. three discs were pasted in each dish.
pH was adjusted to 2.5 with sterile hydrochloric
the distance was more than 24 mm of each
acid. trypsin (sigma) was added into the sim-
disc center and more than 15 mm from disc
ulated pancreatic juices with a final concentra-
edge to the inner edge of dish. next, the dishes
tion of 10 mg/mL. then pH was adjusted to 8.0
were placed at room temperature for 1.5 h and
with 0.1 M naoH.
then incubated at 37°c. After 24 h, the inhibi-
All of the tolerability detection, the initial bac-
tion zone diameter was measured around the
terial counts were adjusted to about 108 cfu/
antibiotic disc with vernier caliper and record-
mL and were checked by viable count determi-
ed. for one tested strain, each antibiotic disc
nation on Mrs agar. for the tolerance assay,
was done 3 times. the inhibition zone diame-
the bacterial suspensions were incubated and
counted at 37°c for 0,1,2,3,4,5,6 h, respectively.
standard sensitive strain of
E. coli Atcc25922
All tests were repeated three times to estimate
was used as the control. the operation was the
the standard error.
same as the above.
the antibiotic susceptibility of the tested
2.5 Detection of antibiotic resistance genes
strains was evaluated according to the cLsI cri-
teria (table 2).
Part of the antibiotic-resistant genes of those
lactic acid bacteria containing both plasmids and
2.3 Plasmid DNA extraction
high tolerance were investigated. the β-lactam
resistance-related gene sequence of
blr,
ECP-
10 mL of
Lactobacillus suspension cultured
1569,
nps-1 and the chloromycetin resistance-
overnight was centrifugated at 10000 rpm for
related gene sequence of
cmlA,
cat,
cmlA1 in
5 min. then the precipitation was suspended
plasmids were found in national center for bi-
with 500 μL of lysozyme solution (10 mg/mL).
otechnology Information (ncbI). the primers
the mixture was placed in a water bath for 45
were designed and synthesized by beijing sun-
min at 37°c. then plasmid dnA of
Lactobacilli
biotech co., Ltd (table 3).
strains was extracted and purified with dnA ex-
the Pcr programmes were performed with the
traction and purification kit of tiangen biotech
plasmid template of the tested strains accord-
(beijing) co., Ltd. Plasmid dnA was observed
ing to the following procedures: initial heating
by agarose gel electrophoresis.
at 94°c for 4 min was followed by 34 cycles of
Antibiotic susceptibility and plasmid stabil-
the following sequence: 94°c for 30 s, 72°c for
ity were tested after cultivated 30 generations
1 min, and 72°c for 1 min. final extension took
at 37°c in Mrs medium according to the above
place at 72°c for 7 min.
the amplification products were separated
284 Ital. J. Food Sci., vol. 27 - 2015
table 3 - the primers of the resistance genes in the experiment.
Sequence of the primer
Annealing temperature
5'-CGTCTTATTGAATTAACAGGTTGG -3'
blr--down
5'-CACGAAGCCATGTTGTGTTC -3'
ECP-1569--up
nps-1--up
nps-1--down 5'-GGCGATACCGCTCAGTTAC-3'
cmlA--down 5'-CATGCCCAAACCTAGAAACGC-3'
cat--down 5'-TGGAAGCCATCACAAACG-3'
cmlA1--up
cmlA1--down 5'-CTACGTTGTGGCGTCAATG-3'
by conventional 1.0% (w/v) agarose gel electro-
ance to 13 kinds of antibiotics. to bacitra-
phoresis (100V, 4°c) in tAE (tris-acetate-Ed-
cin, polymyxin b, kanamycin and nalidixic acid,
tA) buffer and visualised by ethidium bromide
the resistance rate of the 29 tested strains was
staining. the target fragment was recovered
100%. to β-lactam and aminoglycosides, the
and sequenced by taKara biotechnology (dali-
resistance percentage was 20.7%-37.9% and
an, china) co., Ltd. the resistance-related gene
86.2%-100%, respectively. All of the 29 strains
of plasmid was determined by comparison with
were mostly sensitive to chloramphenicol and
Among of the tested antibiotics, the nali-
dixic acid and polymyxin b can inhibite dnA
3. rEsuLts And dIscussIon
synthesis and interfer cell membrane forma-
tion, respectively. the resistance of lactobacil-
3.1 Antibiotic susceptibility
lus to these kinds of antibiotics may be due to
the thicker cell wall of Gram-positive bacteria.
Antibiotic susceptibility of the tested strains
While the tested strains showed different sen-
was evaluated according to the anti-microbi-
sitivity to the antibiotics, such as streptomy-
al drug sensitivity standard of cLsI criteria.
cin, kanamycin, tetracycline, chloramphenicol,
the sensitivity of the tested
Lactobacillus to 13
gentamicin with protein synthesis inhibitition
kinds of antibiotics was shown in table 4. the
effect. Most lactobacillus strains showed re-
tested 29 strains were generally resisitant to
sistance to those antibiotics against gram-neg-
ative bacteria, for example, streptomycin, gen-
lidixic acid, and were mostly sensitive to chlo-
tamicin, kanamycin. this was consistent with
ramphenicol and tetracycline. the same species
report of Zhang
et al (2007).
of
Lactobacillus generally had similar resistance
patterns. but there was species specificity such
3.2 Plasmid DNA extraction
as the different antibiotic sensitivity in
L. plan-
of antibiotics-resistant lactobacillus strains
tarum,
L. rhamnosus, and
L. pentosus. Moreover,
the antibiotic-resistant level of different strains
16 cIcc strains with relatively strong antibi-
is also different.
otic resistance were screened for plasmid extrac-
Antibiotic resistance of the foodborne lactic
tion. As can be seen from fig. 1, among these
acid bacteria had heen reported in the 1980s.
strains, only cIcc 23180, 22161, 22175, 22157,
the researchers generally believed that the re-
23124, and 22154 contained plasmids.
sistance was a result of the long evolution and
L. plantarum cIcc 23180 showed 6 plasmid
it was generally endogenous resistance and ob-
dnA bands and there is one band greater than
tained resistance (Zeng
et al., 2004). so, the re-
23 kb.
L. pentosus cIcc 22157 showed two plas-
sistant lactic acid bacteria of natural or isolat-
mid dnA bands of 10 kb and 5 kb, respective-
ed from human intestinal can indirectly reflect
ly.
L. rhamnosus cIcc 22175 and
L. plantarum
the habitat of used antibiotic.
cIcc 23124 contained respectively 2 and 4 of
It can be seen from table 5, the 29 strains
plasmid dnA bands and both of the two strains
contained a 10 kb plasmid.
L. helveticus cIcc
Ital. J. Food Sci., vol. 27 - 2015 285
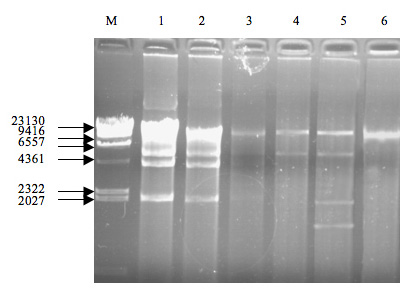
table 5 - the percentage of the antibiotic resistance of 29
Antibiotics
Quantity
of resistant strains
of resistance (%)
penicillin G 11
multi-polymyxin B 29
chloramphenicol 3
L. casei
22154 showed only one plasmid dnA band of
about 10 kb.
L. acidophilus
Lactic acid bacteria generally contain plas-
mids. the plasmid size was usually 1.9 kb-84.8
kb. Most of the plasmid was less than 20 kb
(WAnG and LEE, 1997). In the culture process
from generation to generation, many plasmids
might disappear from the bacterial cell, but most
of the plasmids were stable. In the study, the
plasmids of the above six strains and the anti-
biotic susceptivity showed no changes after cul-
tivated 30 generations.
3.3 Gastrointestinal tolerability
resistance to gastrointestinal stress is very
important for one strain to play the potential
probiotic function (GuGLIELMottI et al., 2007).
If the strains have a high tolerance to gastroin-
testinal stress, it will have the chance to sur-
vive and play the probiotic effects in the gastro-
the tolerance of the selected six strains to low
strains to 13 antibiotics.
fig. 1 - the plasmids in Lactobacillus (1.cIcc 23180, 2.cIcc
22161, 3.cIcc 22175, 4.cIcc 22157, 5.cIcc 23124, 6.cIcc
22154. M. λHindIII marker).
286 Ital. J. Food Sci., vol. 27 - 2015
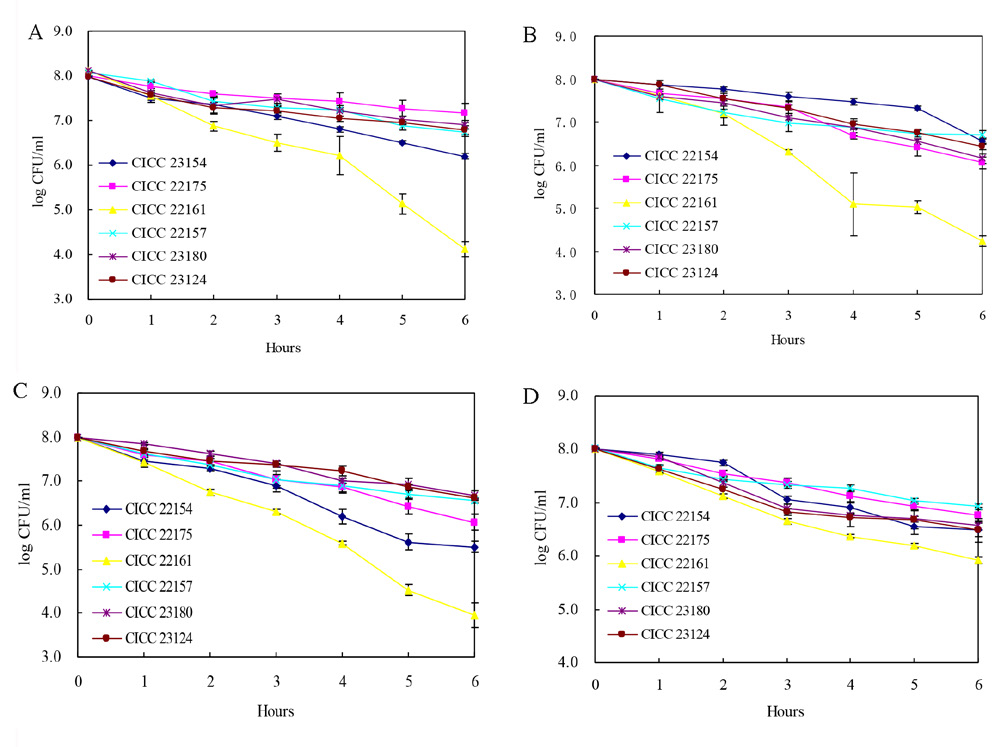
fig. 2 - the viable counts of strains ccIcc 22154, 22175, 22161, 22157, 23180 and 23124 in the gastrointestinal environ-
ment after 6 hs at 37°c.
A: pH 2.5; b: 3 mg/mL bile; c: 5mg/mL pepsin; d: 10 mg/mL trypsin.
pH, bile salt, pepsin and trypsin is presented in
the viable counts of the tested six strains still re-
fig. 2. As shown in fig. 2A, the viable counts of
mained at 106 cfu/mL or more after 6 h exposure
L. pentosus cIcc 22161 strain reduced to be-
to 10 mg/mL trypsin solution (pH 8.0).
low 106 cfu/mL after 4 h and 1.32 × 104 cfu/
mL after 6 h. However, the viable numbers of
3.4 Detection of Resistance genes
L. helveticus cIcc 22154, L. pentosus cIcc
22157, L. plantarum cIcc 23124, 23180 and
According to the above results, except strain
L. rhamnosus cIcc 22175 were still more than
L. pentosus cIcc 22161 and L. helveticus cIcc
106 cfu/mL after 6 h in the gastric acid of pH
22154, the tested strains may be able to sur-
2.5. thus, these five strains showed higher tol-
vive in the simulated gastrointestinal environ-
ment. However, if the above strains contain an-
for bile tolerance, except the L. pentosus cIcc
tibiotics-resistant plasmids, there is the possi-
22161, the viable counts of the other five strains
bility of resistance transfer to other bacteria, es-
were still more than 106 cfu/mL after 6 h in the
pecially pathogenic bacteria. It will be a poten-
medium containing bile salt (fig. 2b). However,
tial hazard to human health and be a serious so-
the viable cells of L. pentosus cIcc 22161 had
cial problem. so, the plasmid-determined resist-
decreased to 2.0×106 cfu/mL within 3 h. And
ant gene should be checked firstly before sub-
it declined to only 1.8 × 104 after 6 h.
for pepsin tolerance, among of six strains,
After 0.02% sds combined with heat treat-
the viable cells of L. pentosus cIcc 22161 and
ment of the four strains (cIcc 22175, 22157,
L. helveticus cIcc 22154 decreased significant-
23124, 23180), only the plasmids of L. plantar-
ly in 6 h and it is less than 104 cfu/mL and 106
um cIcc 23180 were removed and the resist-
cfu/mL after 6 h exposure to 5 mg/mL pepsin
ance to cephalothin and chloromycetin disap-
solution (pH 2.5), respectively (fig. 2c).
peared simultaneously (unpublished results).
for trypsin tolerance, as can be seen in fig. 2d,
so, the primers of β-lactam resistance-relat-
Ital. J. Food Sci., vol. 27 - 2015 287
ed genes including blr, ECP-1569 and nps-1 as
uct types. And studies have shown that more
well as chloromycetin resistance-related genes
genes associated with antibiotic resistance are
including cmlA, cat and cmlA1 were designed.
located in plasmids and transposons (doucEt
the plasmid-determined resistant genes of L.
et al., 1992; MAyyA et al., 2011). but unlike the
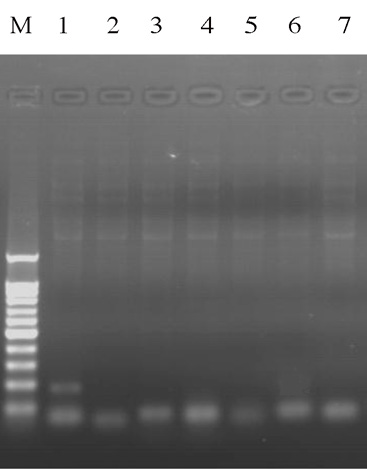
plantarum cIcc 23180 were detected by Pcr.
chromosome dnA, both plasmids and transpo-
As shown in fig. 3, the plasmid of L. plantar-
sons can provide the possibility of transferabili-
um cIcc 23180 contained β - lactam resist-
ty for resistance genes between bacteria. PIEr et
ance gene blr, excluding other resistant genes.
al. (2003) proved the high transferability of plas-
blr gene encodes beta-lactamase, which can
mid pcf10 that encodes tetracycline resistance
hydrolyze β-lactam ring and then make the
from Enterococcus faecalis oG1rf to Enterococ-
β-lactam antibiotic inactivation. this is proba-
cus faecalis bf3098c during cheese and sausage
bly the main reason of the bacteria resistant to
fermentation. JoAnnA et al. (2008) reported the
β-lactam antibiotics. In the present study, the
transferability of erythromycin resistant plasmid
successful amplification of blr gene in L. plan-
(pAMβ1) from Lactococcus lactis sH4174 to Lac-
tarum cIcc 23180 indicated that its cefalotin
tococcus lactis bu2-60. A similar study also indi-
cated that the transferability of tetracycline re-
sistance in E. italicus LMG 22195 from ferment-
ed milk (MIrIAM et al., 2010).
so, the assessment of antibiotic resistant of
potentially probiotic lactic acid bacteria used in
food industry, especially the resistance-relat-
ed genes and the transferability are very neces-
sary. We can also say that, exploring the pro-
biotic property and safety of lactic acid bacte-
ria are equally important.
the tested 29 strains of potential probiotic lac-
tobacillus showed different resistance to antibi-
otics. those resistant strains containing both
plasmids and high tolerance to gastrointesti-
nal condition may cause food safety problems.
so these strains need to be re-assessed carefully.
the study found that the plasmid of L. plantar-
um cIcc 23180 exactly carried the cephaloth-
in-related gene blr. However, the transferibility
of the resistance-related gene remains to be fur-
fig. 3 - the Pcr result in the genome and plasmid of cIcc
ther studied. this study provides a reference in
investigating the relationships between antibi-
M. Marker; 1. blr; 2. EcP-1569; 3. nps; 4. cmlA; 5. cat; 6.
cmlA1; 7.control.
otic resistance spectrum and the plasmids and
evaluating the safety of probiotics.
resistance may be due to the effect of the beta-
lactamase to β-lactam antibiotics.
While in the study, the genes of cat, cmlA and
this work was supported by the science and technolo-
gy research youth fund Project (2010240) and the natu-
cmlA1 were not detected in the plasmids of L.
ral science foundation of Hebei Province (c2013208161,
plantarum cIcc 23180. However, L. plantarum
c2010000863). Authors also thank national High-tech Pro-
cIcc 23180 strain was resistant to chloram-
ject ("863 Plan", no. 2011AA100902) from chinese Ministry
of science and technology for part fund.
phenicol. At the same time, plasmid elimination
and Escherichia coli transformant test showed
that chloramphenicol resistance-related genes
should be present in plasmid dnA of L. plantar-
um cIcc 23180 (unpublished results). there-
charteris W.P., Kelly P.M., Morelli L., collins J.K. 1998a. An-
fore, the plasmid of the cIcc 23180 strain may
tibiotic susceptibility of potentially probiotic Lactobacillus
contain other genes encoding chloramphenicol
species. Journal of food Protection. 61: 1636.
charteris W.P., nelly P.M., Morelli L., collins J.K. 1998b.
development of an in vitro methodology to determine the
In recent years, more studies have been done
transit tolerance of potentially probiotic Lactobacillus and
on antibiotic resistance of probiotics. It was
bifidobacterium species in the upper human gastroin-
shown that the antibiotic resistance was varia-
testinal tract. Journal of Applied bacteriology. 84: 759.
ble, species-dependent and related to the prod-
doucet P.f., trieu c.P., Andremont A., courvalin P. 1992.
288 Ital. J. Food Sci., vol. 27 - 2015
conjugal transfer of plasmid dnA from Enterococcus fae-
Liu y.Q., Zhou f., Zhao H.f., Zhan H.n., Zhang b.L. 2011.
calis to Escherichia coli in digestive tracts of gnotobiotic
factors affecting the production of folic acid by lactic acid
mice. Antimicrobial Agents and chemotherapy. 36(2): 502.
bacteria. china dairy Industry. 39(3): 10.
franz c.M.A.P., Hummel A.P., Holzapfel W.H. 2005. Prob-
Maria r.d., Monica M., bruno b. 2007. Antibiotic resistance
lems related to the safety assessment of lactic acid bacte-
of lactic acid bacteria and bifidobacterium spp. isolated
ria starter cultures and pr
from dairy and pharmaceutical products. International
Journal of food Microbiology. 115: 35.
Guglielmotti d.M., Marco´ M.b., Golowczyb M., treinherim-
Mayya P., Zhosephine G., sofia M. 2011. tn5045, a novel
er J.A., Quiberoni A.L. 2007. Probiotic potential of Lac-
integron-containing antibiotic and chromate resistance
tobacillus delbrueckii strains and their phage resistant
transposon isolated from a permafrost bacterium. re-
mutants. International dairy Journal. 17: 916.
search in Microbiology. 162: 337.
Hummel A., Holzapfel W.H., franz c.M. 2007. character-
Miriam Z., Geert H., Giorgio G. 2010. Molecular basis and
isation and transfer of antibiotic resistance genes from
transferability of tetracycline resistance in Enterococcus
enterococci isolated from food. systematic and Applied
italicus LMG 22195 from fermented milk. International
Microbiology. 30: 1.
Journal of food Microbiology. 142: 234.
Huys G., d'Haen K.d., collard J.M., swings J. 2004. Prev-
niamh t., declan b., se´amus f. 2010. characterisation and
alence and molecular characterization of tetracycline re-
transferability of antibiotic resistance genes from lactic
sistance in Enterococcus isolates from food. Applied and
acid bacteria isolated from Irish pork and beef abattoirs.
Environmental Microbiology. 70: 1555.
research in Microbiology. 161(2): 127.
Jin s., Zhang G.L., Ji d.d., Zhang b.L. 2009. study on lac-
Pier s.c., daniela c., simona G. 2003 Gene transfer of van-
tic acid bacteria on inhibiting mutagenic and carcino-
comycin and tetracycline resistances among Enterococcus
genic substances. science and technology of food In-
faecalis during cheese and sausage fermentations. Inter-
dustry. 30(12): 165.
national Journal of food Microbiology. 88: 315.
Joanna L., Louise f., Aine M., niamh t., susanne s., bod-
sun X.Q., Zhang X.L., Wang s., Zhang b.L. 2009. optimized
il J., Hilko van der Voet, sigrid r.A., declan b., Henk A.,
production and application of EPs by Lactobacillus pen-
Karen A.K., Andrea W., Jacek b. 2008. A standardized
tosus strains in fermented milks. Journal of dairy sci-
conjugation protocol to asses antibiotic resistance trans-
ence and technology. 5: 212.
fer between Lactococcus species. International Journal of
threlfall E.J., Ward L.r., frost J.A., Willshaw G.A. 2000.
food Microbiology. 127: 172.
the emergence and spread of antibiotic resistance in
food-borne bacteria. International Journal of food Mi-
crobiology. 62: 1.
s., H. 2007. Antimi-
Wang t.f., Lee b.H. 1997. Plasmids in Lactobacillus. criti-
crobial susceptibilities of Lactobacillus, Pediococcus and
cal reviews in biotechnology. 17(3): 227.
Lactococcus human isolates and cultures intended for
probiotic or nutritional use. Journal of Antimicrobiology
hite d.G., Zhao s., simjee s., Wagner d.d., Mcdermott P.f.
chemotherapy. 59: 900.
2002. Antimicrobial resistance of foodboe pathogens. Mi-
crobes and Infection. 4: 405.
lection and maintenance of probiotic microorganisms.
Zeng H.y., Qin L.K., Jiang P. 2004. development review on
In: Lee, y.K. and salminen, s. (2nd, Ed.), Handbook of
acquired antibiotic resistance in lactic acid bacteria from
probiotics. John Wiley & sons, new york, pp 177-188.
food. food science. 25(12): 189.
Li sh.y., Li .Pf., shi J.H., Lei sh.ch., Zhang y.y., Zhang
Zhang Z.y., Liu c., Guo X.K. 2007. research progress of an-
K.P. 2008. Isolations of the bifidobacterium from cows
tibiotics resistance in lactic acid bacteria. chinese Jour-
and their resistance characteristics to given antibacteri-
nal of Microecology. 19(5): 478.
al drugs. dairy Industry china. 1: 1.
Li ch., Wang s., Zhan H.n., Zhao H.f., Pei J.W., Zhang
screening of Lactobacillus strains
b.L. 2009. roles of Lactobacillus paralimentarius 412 in
for their ability to bind benzo(a)pyrene and the mech-
sourdough fermentation. food and fermentation Indus-
tries. 35(5): 99.
Paper Received January 16, 2014 Accepted August 25, 2014
Ital. J. Food Sci., vol. 27 - 2015 289
Source: http://www.chiriottieditori.it/ojs/index.php/ijfs/article/download/270/41
Dr.med. Josef Zehentbauer (b.1945) - www.josef-zehentbauer.de/ - Arzt und Psychotherapeut in München Wer hat Angst vor dem Psychopharmaka Absetzen? Ä r z t l i c he B er at ung und p sy c hot her ape ut i sc he G espr äc he bei m A bset z en v on Däm pf ungs - und B er u hi gun g sm i tt el n Neuroleptika / Antidepressiva / Phasenprophylaktika: Carbamazepin, Lithium / Tranquilizer
Sistema de Composición Corporal Fit Fit Reset Clnsng Bev Tropical Shrnk Slv 95% de todas las dietas fallan. A quí está tu oportunidad de ser parte de un 5 por ciento cool gray 7 pms 1505 pms 173 .white Fit es un enfoque inteligente y saludable para la perdida de grasa al combinar poderosos bioactivos a base de productos con una nutrición balanceada y con ejercicio. Esta científicamente diseñado y probado para ayudar a optimizar tu composición corporal mediante la limpieza de tu cuerpo mientras



