Kamagra gibt es auch als Kautabletten, die sich schneller auflösen als normale Pillen. Manche Patienten empfinden das als angenehmer. Wer sich informieren will, findet Hinweise unter kamagra kautabletten.
Pc131086 1.20
This article is a Plant Cell Advance Online Publication. The date of its first appearance online is the official date of publication. The article has beenedited and the authors have corrected proofs, but minor changes could be made before the final version is published. Posting this version onlinereduces the time to publication by several weeks.
A Secreted Effector Protein of Ustilago maydis Guides MaizeLeaf Cells to Form Tumors
Amey Redkar,a,1 Rafal Hoser,b Lena Schilling,a Bernd Zechmann,c Magdalena Krzymowska,bVirginia Walbot,d and Gunther Doehlemanna,e,2
a Max Planck Institute for Terrestrial Microbiology, Department of Organismic Interactions, D-35043 Marburg, Germanyb Institute of Biochemistry and Biophysics, Polish Academy of Sciences, 02-106 Warsaw, Polandc Baylor University, Center for Microscopy and Imaging, Waco, Texas 76798d Department of Biology, Stanford University, Stanford, California 94305e Botanical Institute and Cluster of Excellence on Plant Sciences, University of Cologne, 50674 Cologne, Germany
The biotrophic smut fungus Ustilago maydis infects all aerial organs of maize (Zea mays) and induces tumors in the plant tissues.
U. maydis deploys many effector proteins to manipulate its host. Previously, deletion analysis demonstrated that several effectorshave important functions in inducing tumor expansion specifically in maize leaves. Here, we present the functional characterizationof the effector See1 (Seedling efficient effector1). See1 is required for the reactivation of plant DNA synthesis, which is crucial fortumor progression in leaf cells. By contrast, See1 does not affect tumor formation in immature tassel floral tissues, where maize cellproliferation occurs independent of fungal infection. See1 interacts with a maize homolog of SGT1 (Suppressor of G2 allele of skp1),a factor acting in cell cycle progression in yeast (Saccharomyces cerevisiae) and an important component of plant and humaninnate immunity. See1 interferes with the MAPK-triggered phosphorylation of maize SGT1 at a monocot-specific phosphorylationsite. We propose that See1 interferes with SGT1 activity, resulting in both modulation of immune responses and reactivation of DNAsynthesis in leaf cells. This identifies See1 as a fungal effector that directly and specifically contributes to the formation of leaftumors in maize.
liberating a sooty mass of black teliospores (Brefort et al., 2009).
Among Ustilaginales, Ustilago maydis, a model organism for bio-
To establish a successful infection and dampen plant defense
trophic fungi (Kämper et al., 2006; Ökmen and Doehlemann, 2014),
responses during colonization, plant pathogens secrete proteins
has the unique ability to colonize all the aerial organs of its host
and other molecules, collectively termed effectors, to various
plant maize (Zea mays) and to induce the formation of plant tumors
host compartments (Jones and Dangl, 2006). Effectors are key
locally at sites of infection. The fungus penetrates the epidermal
to the alterations of host structures and functions during infection
cells and then the subepidermal cells, forming an interaction zone
(Hogenhout et al., 2009). They act either in the intercellular space
called the biotrophic interface in which the hyphae are encapsu-
to handle the primary defense response or inside the host cell to
lated by the host plasma membrane. After successful establish-
execute functions such as reprogramming of the host to favor
ment in leaves, the fungus grows in the mesophyll and the living
infection (Doehlemann et al., 2014).
cells of the vasculature (Ökmen and Doehlemann, 2014). Prolifer-
The basidiomycetous plant pathogens are highly specialized
ation of host and fungal cells results in tumors, which are supported
colonizers that develop biotrophic interactions. Members of the
by the comprehensive reprogramming of both plant signaling and
Ustilaginales, a major order of this class, invade mainly monocots,
metabolism early in infection (Doehlemann et al., 2008; Horst et al.,
including all major cereal crops. The infection normally occurs in
2010) and alteration of the pace and pattern of host cell division.
seedlings, often immediately after mating of the compatible spor-
The U. maydis genome encodes ;550 proteins that are pre-
idia to form a dikaryotic filament (Kämper et al., 2006; Brefort et al.,
dicted to be secreted and likely function as effectors (Mueller et al.,
2009). Fungal hyphae growing both intracellularly and intercellu-
2008; Djamei and Kahmann, 2012). Many potential effector genes
larly colonize the host systemically and grow toward the shoot
are arranged in clusters, and examination of deletion mutants re-
apical meristem without inducing visible disease symptoms. Dis-
vealed the importance of these genes in virulence (Kämper et al.,
ease symptoms become evident upon the floral transition, and the
2006; Brefort et al., 2014). So far, only a few effector genes of
fungus completes sporulation within the infected inflorescence,
U. maydis have been functionally characterized. Pep1 (Proteinessential for penetration1) is involved in penetration and the es-
tablishment of initial compatibility by targeting and inhibiting the
Current address: The Sainsbury Laboratory, Norwich Research Park,
Norwich NR47UH, UK.
activity of the plant peroxidase POX12 (Doehlemann et al., 2009;
2 Address correspondence to
Hemetsberger et al., 2012). Pit2 (Protein involved in tumors2), a
The author responsible for distribution of materials integral to the findings
protein essential for tissue colonization and plant defense suppres-
presented in this article in accordance with the policy described in the
sion, inhibits apoplastic cysteine proteases (Doehlemann et al., 2011;
Instructions for Authors (is: Gunther Doehlemann
Mueller et al., 2013). In addition, two translocated U. maydis ef-
fectors have been analyzed. The U. maydis chorismate mutase
The Plant Cell Preview, www.aspb.org ã 2015 American Society of Plant Biologists. All rights reserved.
Cmu1 rechannels chorismate metabolism in the plant cell cyto-
both maize inflorescences and aerial vegetative tissues, such as
plasm to prevent the synthesis of salicylic acid, a major defense
seedling leaves (Skibbe et al., 2010). A previous analysis of
signal (Djamei et al., 2011). The effector Tin2 (Tumor inducing2),
U. maydis effector candidates with organ-specific expression
which is part of the largest cluster of effectors in U. maydis
patterns identified seven genes whose deletion resulted in a leaf-
(Brefort et al., 2014), masks a ubiquitin-proteasome degradation
specific reduction of tumor formation (Schilling et al., 2014), and
motif in TTK1, a maize protein kinase that regulates the antho-
here we investigate one of these genes (um02239, now termed
cyanin biosynthetic pathway. Tin2 protects the active kinase
see1) for its specific role.
against ubiquitination and thereby promotes the production of
Deletion mutants of see1 (SG200Δsee1) mainly formed tumors
anthocyanin in infected tissue and suppresses lignin biosyn-
of 1 to 4 mm in diameter on seedling leaves at 12 DPI; these
thesis, a defense pathway (Tanaka et al., 2014).
symptoms represent about half of the total tumors formed. Tu-
U. maydis infects all maize aerial organs and thus interacts
mors of >6 to 20 mm occurred frequently in wild-type infections,
with different, developmentally distinct immature host tissues
representing around 28% of the total tumors, but they occurred
(Walbot and Skibbe, 2010). In a previous study, organ-specific
much less frequently and were reduced in size in SG200Δsee1
transcriptomes of both the host and the pathogen were docu-
infections, representing only 9% of tumors. Heavy tumors, which
mented in seedlings, adult leaves, and tassels (Skibbe et al., 2010).
cause altered leaf shape or even stunted growth of infected
It was hypothesized that effectors in U. maydis act in an organ-
seedlings, were not observed after infection by SG200Δsee1 (Figure
specific manner, a new concept now extended to anthers within
1A; ). The SG200Δsee1 mutant induces
the tassels (Gao et al., 2013). A recent study showed that individual
normal tumors in maize tassels, indistinguishable from the virulent
effector genes of U. maydis act in specific plant organs and that
progenitor strain SG200 (Figures 1A and 1B). Teliospores dissected
deletion of one organ-specific effector does not hamper virulence
from these tassel tumors were normal in shape and fully viable
in a nontarget organ (Schilling et al., 2014). To date, however, the
). Similarly, in the maize ear, tumor formation
functional basis of organ-specific effectors remains elusive.
was comparable to that in the wild type, supporting a strictly leaf-
Effectors may be recognized by plant receptor proteins, which
specific role of See1 in tumor induction .
in turn induce defense responses. Several plant receptor proteins
Confocal microscopy showed that SG200Δsee1 hyphae initially
function with the help of chaperones and cochaperones, in-
colonize similarly to the progenitor strain SG200. At 3 DPI, when
cluding HSP90 (heat shock protein 90), RAR1 (required for Mla12
wild-type fungal hyphae reach the leaf mesophyll and are inter-
resistance), and SGT1 (suppressor of G2 allele of skp1) (Shirasu,
spersed within the vasculature, mutant hyphae clustered at col-
2009; Zhang et al., 2010). SGT1 was originally identified in Sac-
lapsed, highly fluorescent mesophyll cells
charomyces cerevisiae as an essential cell cycle protein that in-
). In addition, mutant hyphae failed to traverse from an infected
teracts with Skp1p, a component of the conserved eukaryotic
cell into uninfected neighboring cells; this was particularly observed
Skp1/Cullin/F-box (SCF) E3 ubiquitin ligase. In yeast, Sgt1p is
in bundle-sheath cells Reintroduction of
required for progression through the G1/S and G2/M checkpoints
the see1 gene into the U. maydis ip locus fully restored virulence,
(Kitagawa et al., 1999) and is highly conserved, as its orthologs in
demonstrating functional complementation of see1 and confirming
both animal and plant kingdoms retain the cell cycle functions
that the observed growth defects reflected the absence of See1
(Bhavsar et al., 2013). Maturation of SGT1 as a signaling molecule
(Figures 1A and 1B).
depends on phosphorylation by an upstream MAPK (Hoser et al.,
Transcription of see1 specifically increased during biotrophic
growth of U. maydis (Figure 2A). Comparison of the temporal
In this study, we present the functional characterization of
and spatial profiles of see1 expression during successive stages
the U. maydis organ-specific effector See1 (Seedling efficient
of tumor progression showed that see1 expression is constitu-
effector1; Um02239), which is specifically required during tumor
tive and then upregulated at the later stages of tumor expansion
formation in seedling leaves. See1 is translocated by the fungus
in maize leaves but not in tassels (Figure 2A). In maize ear tumors,
into the plant cell cytoplasm and nucleus, where it interacts with
see1 transcript abundance was low as in tassels at 12 DPI. At
the maize homolog of SGT1 and interferes with the MAPK-
this time point, see1 expression was >50-fold induced in leaves
induced phosphorylation of SGT1. See1 participates in U. maydis-
compared with the floral organs To gain
triggered reactivation of plant DNA synthesis in maize leaves and
comprehensive insight into host processes affected by see1 de-
contributes to vegetative tumor formation.
letion, Agilent microarrays were used to profile the transcriptomeof maize leaves at 6 DPI by SG200, SG200Δsee1, and mockcontrol infections. RNA of infected tissue was prepared from three
biological replicates, analyzed by hybridization, and subjectedto data normalization and statistical analysis (see Methods fordetails). The abundances of 10,952 maize transcripts were al-
See1 Is Required for the Induction of Leaf Tumors
tered in response to infection with wild-type U. maydis; by con-
After infection, U. maydis hyphae mainly grow intracellularly.
trast, only 773 transcripts were altered in response to infection
About 4 d postinfection (DPI), small tumors are visible and the
with SG200Δsee1 and
fungus proliferates massively both intracellularly and intercellu-
). Hierarchical clustering of the SG200-induced maize genes
larly. In mature tumors at 10 to 14 DPI, U. maydis forms masses
visualized the reduced transcriptional response of maize leaves to
of melanized teliospores (Doehlemann et al., 2008). Unlike other
the see1 deletion mutant . A direct com-
smut fungi of monocots, U. maydis causes these symptoms in
parison of SG200 with SG200Δsee1 showed that 549 genes were
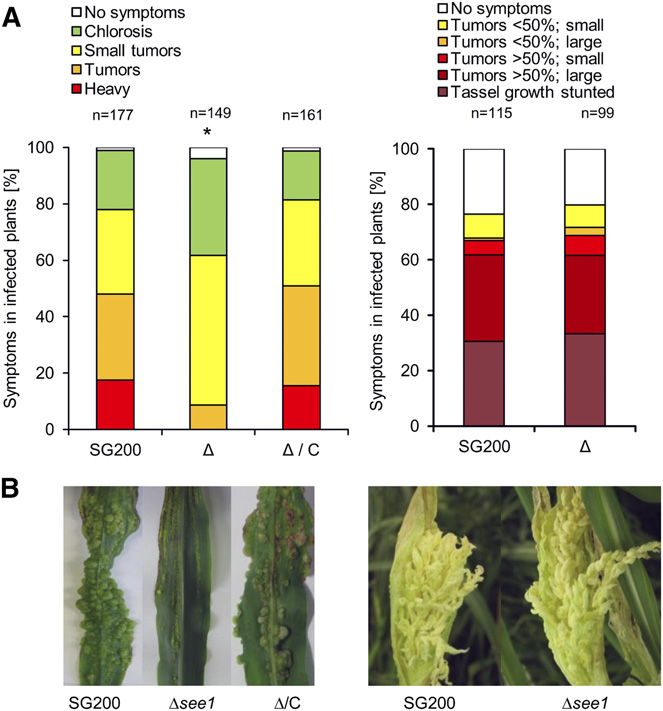
Tumor Induction by Ustilago maydis
regulation of cell division (TC307447) increased 230-fold in wild-type infections . Together, these datasuggest that SG200Δsee1 fails to induce leaf tumor growth at thelevel of host cell DNA synthesis and cell proliferation, processesthat are hallmarks of maize responses to infection (Doehlemannet al., 2008).
See1 Is Required for U. maydis-Induced Plant DNASynthesis during Leaf Tumor Formation
Because see1 expression is prominent during tumor enlargementand our initial observations indicated that SG200Δsee1 hyphaewere mainly restricted within mesophyll and bundle-sheath cells,we performed a more thorough confocal microscopy investigationof leaf infections. U. maydis-induced tumor growth reflects hostproliferation, then cell expansion; thus, DNA synthesis is a pre-requisite for growth. To monitor DNA synthesis in planta, wetreated uninfected and infected leaves with 5-ethynyl-2-deoxy-uridine (EdU) at several time points over a period of 5 h and thenharvested samples. Incorporation of EdU was visualized by at-taching a fluorescent tag (AF-488). Maize nuclei were stained withpropidium iodide (PI) following a procedure described previouslyfor maize anthers (Kelliher and Walbot, 2011). In maize leaves at2 DPI, EdU treatment did not result in any detectable labeling. Weobserved this in maize leaves colonized with U. maydis and in
Figure 1. Organ-Specific Phenotype of See1 Demonstrating Its Role in
uninfected maize leaves (Figure 3A), suggesting that no or only
rare, sporadic maize DNA synthesis occurs in seedling leaf blades
(A) Disease symptoms caused by SG200Δsee1 in comparison with the
during the early phase of infection. We conclude that in the in-
wild-type progenitor strain SG200 in leaves and tassels. The mutant
fected zones, the host cells were already postmitotic. By 4 DPI,
shows a significant reduction in leaf virulence. Maize seedling leaves were
when the first macroscopic symptoms appear in wild-type in-
scored at 12 DPI. Disease symptoms in maize tassels were scored at 14
fections, EdU incorporation into leaf DNA was widespread (Figure
DPI as described by Schilling et al. (2014). SG200, the virulent U. maydis
3A). Leaf cells invaded by fungal hyphae synthesized new DNA,
progenitor strain; D, deletion mutant for see1; D/C, genetic complemen-tation of the deletion strain. The experiment was performed in three in-
and this coincided with the induction of mitosis, which could be
dependent biological replicates. n = number of plants infected. *P # 0.001.
visualized at different stages of cell division and as contiguous
(B) Symptoms caused by U. maydis strain SG200 in comparison with the
pairs of similarly labeled cells (Figure 3B). Such invaded cells also
SG200Δsee1 mutant and the complemented strain in leaves and tassels.
underwent multiple division events over several days (Figure 3B).
The leaf photograph shows typical disease symptoms at 12 DPI; the
As an additional negative control for the EdU labeling, we
tassel photograph is at 14 DPI. Similar to SG200, the mutant caused
injected 5 mM hydroxyurea, a DNA synthesis inhibitor, into
disease symptoms in tassels, but leaf tumors were significantly reduced.
seedling leaves infected with wild-type SG200 or SG200Dsee11 d before labeling them with EdU. Pretreatment with hydroxy-
significantly induced (>2-fold) in SG200 compared with SG200Δsee1
urea eliminated EdU incorporation in all samples; this was also
at 6 DPI, while only two genes were repressed
true in SG200-infected cells that already initiated division before
The transcripts induced by infection with wild-type
treatment. Hydroxyurea appears to block DNA synthesis com-
pathogen were enriched for genes involved in DNA modification
pletely and validates the specificity of the EdU labeling assay
(i.e., histones), DNA replication, and DNA damage repair as well
Quantification of EdU labeling showed
as genes associated with the cell cycle (
that 67.5% 6 4.2% of the maize cells colonized by SG200 in-
. Gene Ontology (GO) analysis showed that 71 of the 549
corporated EdU at 4 DPI (Figures 4A and 4C). Labeling was
SG200-induced genes are associated with DNA metabolism and
initiated from 3 to 4 DPI, while no DNA synthesis was observed
cell cycle regulation (and
in uninfected leaves of the same age. Therefore, U. maydis re-
list the top 30 GO terms that are associated with DNA
activated DNA synthesis and cell division in maize leaves at the
metabolism and cell cycle regulation). As shown in Figure 2B,
onset of tumor induction. By contrast, SG200Δsee1-infected
DNA replicase D (TC280511), which is involved in S-phase DNA
leaf samples showed only 7.3% 6 1.7% EdU-positive cells at
replication, was induced 690-fold in wild-type infections com-
4 DPI (Figures 4A and 4C; The
pared with SG200Δsee1. DNA histone H3 (TC298222), which is
SG200Δsee1 deletion mutant fails to trigger DNA synthesis and
required to generate nucleosomes, was induced 862-fold in wild-
cell division to support the formation of large tumors.
type infections. Maize Skp1 (TC293032) was induced 875-fold in
One might argue that the reduction in host cell DNA synthesis
SG200 infections versus SG200Δsee1. Also, a Leu-rich repeat
could be a general consequence of reduced virulence (i.e.,
receptor-like protein responsible for protein phosphorylation and
an impaired biotrophic interaction); therefore, it would not be
observed (Figure 4A). This observation indicates that biotrophiccolonization of maize smut per se does not induce host DNAsynthesis. Reactivation of DNA synthesis is specific to tumorformation. Next, a U. maydis deletion mutant lacking the secretedeffector Tin3 was tested. Deletion of tin3 results in small leaf tu-mors similar to SG200Δsee1 strains (Brefort et al., 2014). Despite
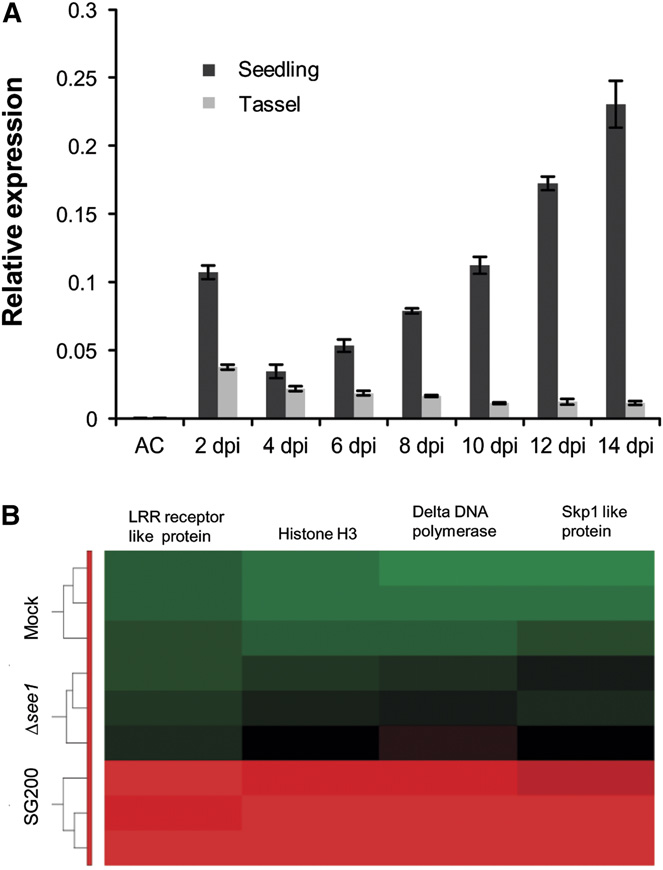
Figure 2. Gene Expression during Maize Colonization with SG200 andSG200Δsee1.
(A) RT-qPCR expression profiling of the see1 gene during the biotrophicphase of U. maydis growth in seedling and tassel tissues. Expressionlevels are shown relative to mean expression of ppi transcripts. Geneexpression was analyzed in axenic culture (AC), seedling, and tasseltissues at consecutive time points from 2 to 14 DPI. The experiment wasperformed in three independent biological replicates.
(B) Transcriptional regulation of the key genes involved in the process ofDNA synthesis and histone modification between wild-type SG200- andSG200Dsee1 (mutant)-infected seedlings at 6 DPI. Hierarchical cluster-ing was performed by the Partek Genomics Suite version 6.12 to visu-alize the expression of maize genes transcriptionally regulated at 6 DPI
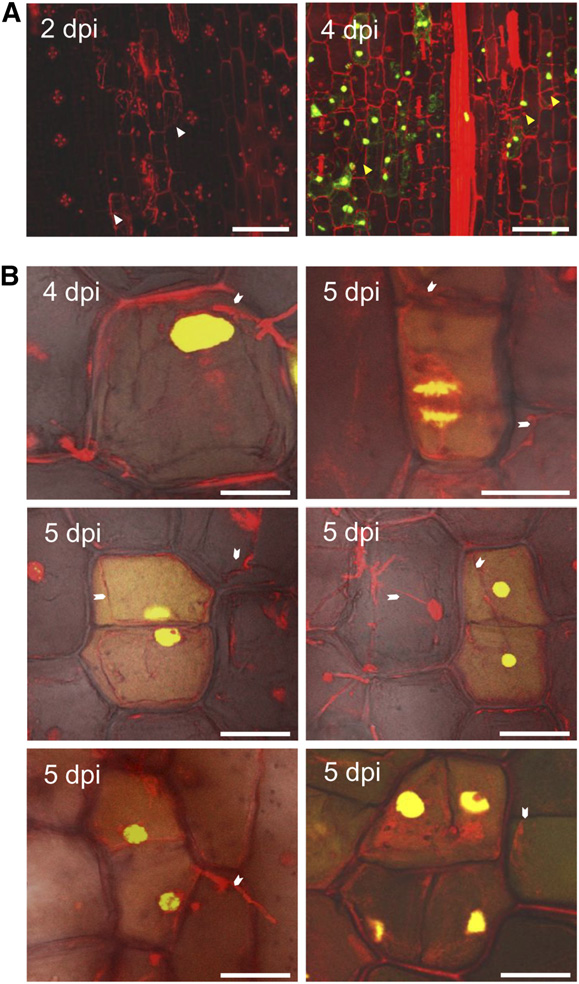
Figure 3. U. maydis Induces DNA Synthesis in Infected Maize Seedlings.
by U. maydis strain SG200 (bottom), infection by SG200Dsee1 (middle),and mock inoculation (top). The x axis depicts clustering of the micro-
(A) Maize seedlings were infected by U. maydis wild-type strain SG200,
array samples for each of the three biological replicates for each treat-
and then tissue was incubated in EdU to visualize in vivo DNA synthesis
ment. The y axis shows clustering of the regulated maize transcripts
in the host cells. Samples were imaged at 2 and 4 DPI by confocal mi-
based on the similarity of their expression patterns. red, upregulated
croscopy. Left, at 2 DPI, the fungal proliferation was observed sub-
genes; green, downregulated genes; black, not significantly altered. LRR,
epidermally; host cells adjacent to fungal hyphae were considered to be
Leu-rich repeat.
colonized cells (white arrowheads). No EdU incorporation was observed.
Right, at 4 DPI, numerous colonized cells showed EdU labeling (greenstain), indicating the onset of DNA synthesis in host cells (yellow ar-
functionally linked with the action of See1. To test this, we in-
rowheads). Bars = 75 mm.
cluded two additional strains. First, leaves were infected with wild-
(B) Cell division events were observed in maize seedlings infected byU. maydis wild-type strain SG200 at 4 and 5 DPI. EdU incorporation into
type strains of the maize head smut fungus Sporisorium reilianum,
a cell will result in equally labeled contiguous daughter cells after cell
a close relative of U. maydis. S. reilianum also establishes a bio-
division. Such equally labeled cell pairs were readily observed in SG200-
trophic interaction with maize, but it causes visible symptoms only
infected seedling leaf tissue. The white arrowheads point to fungal hy-
in the inflorescences, never in leaves (Schirawski et al., 2010).
phae associated with maize cells undergoing cell division. It is inferred
Strikingly, at 4 DPI, the leaves infected with S. reilianum lacked
that reactivation of the cell cycle and rapid divisions are responsible for
detectable DNA synthesis, although dense tissue colonization was
tumor formation. Bars = 25 mm.
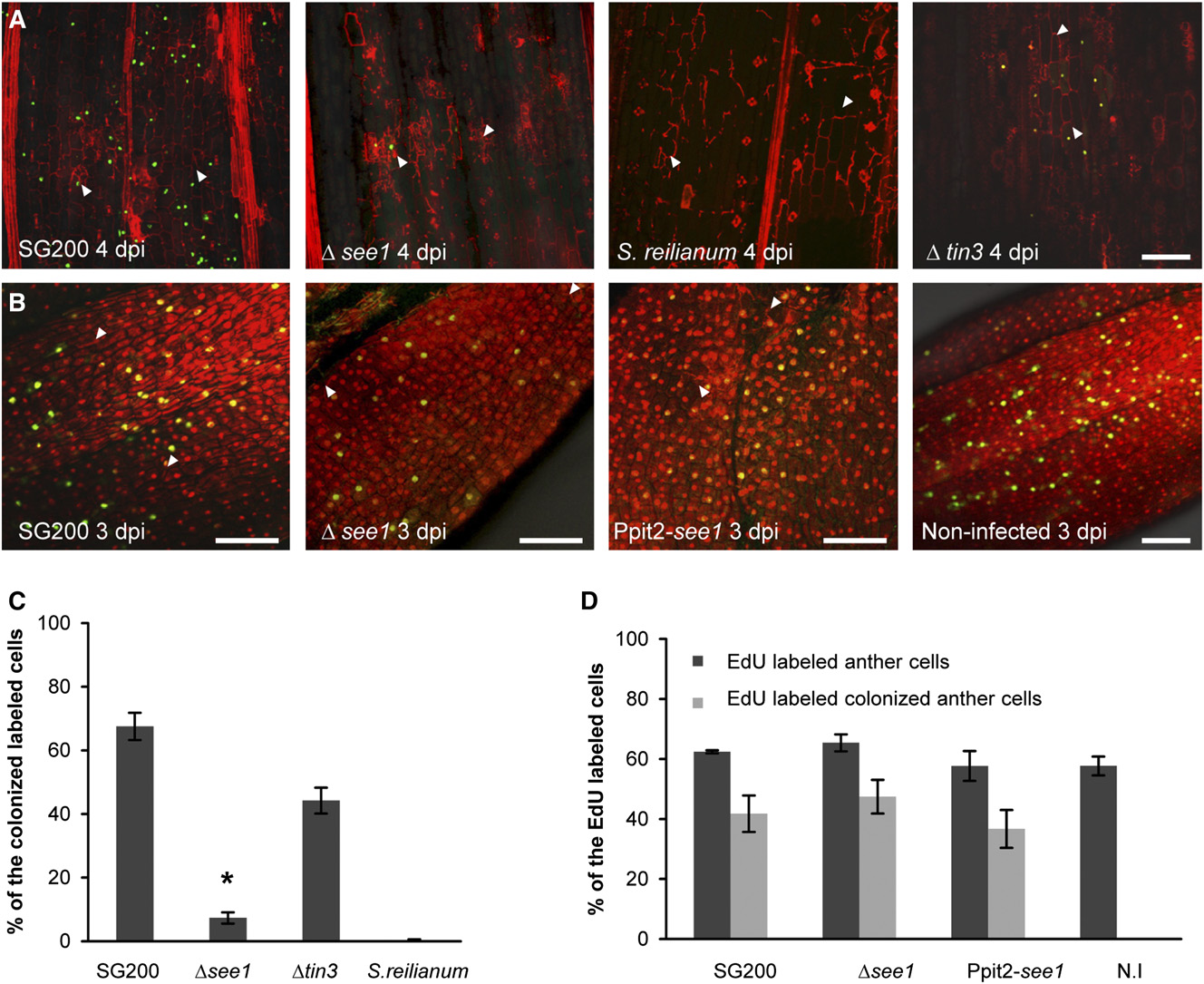
Tumor Induction by Ustilago maydis
Figure 4. See1 Requirement for Host Cell Cycle Release in Leaf Tumor Formation.
(A) In vivo DNA synthesis in seedling tissue infected with SG200Δsee1 in comparison with wild-type SG200. Samples infected with S. reilianum andSG200Δtin3, which has a similar phenotype to SG200Δsee1 with respect to tumor size, were used as controls. Fungal hyphae and plant cell walls werevisualized by PI staining (red), and the EdU-labeled host cell nuclei are visualized by AF488 staining (green). Fungal hyphae are shown by the whitearrowheads. Bar = 100 mm.
(B) DNA synthesis in anther tissue infected with SG200Δsee1 in comparison with wild-type SG200. Samples infected with the strain overexpressingSee1 and uninfected anthers served as controls (right panel). Nuclei were visualized by PI staining (red), and EdU-labeled host cell nuclei are visualizedby AF488 staining (green). Fungal hyphae are marked by white arrowheads. Bars = 100 mm.
(C) Quantification of the EdU-labeled seedling leaf cells in the in vivo DNA synthesis assay comparing infections with wild-type SG200, SG200Δsee1,SG200Δtin3, and S. reilianum. Error bars show SE. *P # 0.001.
(D) Quantification of the EdU-labeled nuclei relative to total anther nuclei per image examined after infection with wild-type SG200, SG200Δsee1, See1-overexpressing strain Ppit2-see1, and noninfected (N.I) tissue. Within the population of EdU-positive cells, the number colonized by fungal hyphae wasalso quantified in the infected samples. Error bars show SE.
its severely reduced virulence, the SG200Δtin3 deletion mutant
indirect consequence of reduced tumor size but reflects a required
activated EdU labeling in 44.22% 6 4.0% of colonized leaf cells at
activity of the See1 effector.
4 DPI (Figures 4A and 4C). Therefore, there is more than onecause of impaired tumor induction, separating See1 from other
Tumor Formation in Anthers Does Not Involve
mutants that lack large tumors but retain the ability to reactivate
U. maydis-Induced DNA Synthesis
widespread host DNA synthesis. From these results, we concludethat the inability of the SG200Δsee1 mutant to reactivate maize
The reproductive spikelets each contain two florets with three an-
cell DNA synthesis and proliferation for tumor formation is not an
thers and arise within the tassel inflorescence; the nonreproductive
floral tissues such as the glumes, palea, and lemma of each spikelet,
with SG200, SG200Δsee1, and noninfected samples (Figures 4B
as well as the tassel stem, are readily infected and transformed to
and 4D), further evidence that See1 is not involved in tumor for-
tumors by U. maydis. Interestingly, the fungus is only effective in
mation in anthers. In the terminal node/internodes, however, the
causing anther tumors during the period of rapid anther growth by
frequency of EdU-labeled cells was significantly increased by see1
cell division prior to meiosis (Walbot and Skibbe, 2010). The intrinsic
overexpression as compared with SG200 (Ppit2-see1, 31.8% 6
anther developmental program of rapid proliferation is reprogram-
4.5%; SG200, 20.7% 6 4.5%; mock, 8.1% 6 1.6%). Therefore,
med into a tumor pathway, with different cell types affected de-
the abnormal phenotype is a direct consequence of the excessive
pending on when fungal hyphae invade the cells (Gao et al., 2013).
cell division resulting from see1 overexpression (Figures 5C and
From this observation, Gao et al. (2013) concluded that tumor for-
5D). In summary, we conclude that See1 is required to stimulate
mation in anthers mainly occurs by restructuring of the usual se-
U. maydis-induced tumor formation by promoting host DNA
quential events in cell fate specification. In line with this hypothesis,
synthesis in vegetative tissue but not in maize anthers.
we did not observe significant differences in EdU-labeled cells inuninfected tissue compared with U. maydis-infected anther tissue.
See1 Localizes to Maize Cytoplasm and Nuclei
Uninfected anthers as well as SG200- and SG200Δsee1-infectedpremeiotic anthers contained ;60% cells labeled with EdU in a 5-h
Live cell imaging and immunolabeling using transmission elec-
treatment (Figures 4B and 4D). This is consistent with the previous
tron microscopy (TEM) were used to localize See1 in planta. An
report that, during the rapid proliferation period of anthers, EdU in-
mCherry-tagged version of See1 lacking its N-terminal secretion
corporation was found in the majority of cells after a 4-h labeling
signal (35S promoter:See1
-mCherry) was transiently expres-
(Kelliher and Walbot, 2011). Therefore, in contrast with leaves,
sed in maize leaves by particle bombardment (Figure 6A).
U. maydis does not alter anther cell DNA synthesis. In line with
-mCherry localized to both the maize cytoplasm and
this, ;42% of the EdU-positive anther cells were colonized by U.
nuclei (Figure 6A). As a transformation control, the nuclear marker
maydis at 4 DPI, with no significant difference between SG200 and
protein PCNA-interacting protein (PIP
)-yellow fluorescent
SG200Δsee1. We conclude that See1 is not involved in modulating
protein (YFP) was coexpressed and localized exclusively to the
host DNA synthesis and cell division during colonization and tumor
nuclei. As a localization control, mCherry expressed alone and
induction in anthers and that it is dispensable for tumor formation in
Pit2-mCherry (Pit2
-mCherry) also showed the same locali-
zation pattern to the cytoplasm and nucleus when transientlyexpressed in maize epidermal cells ). Interestingly, the See1
-mCherry signal spread to
See1 Actively Contributes to Tumor Formation and Maize
the nuclei of cells surrounding individual transformed cells (Figure
6A; ). Fluorescence signal movement
To test whether See1 actively contributes to tumor formation
was not observed for mCherry or for the Pit2-mCherry fusion
and DNA synthesis, a U. maydis strain was generated that con-
protein (. These results in-
stitutively expresses see1 during the entire infection process, in-
dicate that See1 may traffic in planta.
dependently of the colonized tissue. See1 was expressed from the
In addition to the heterologous expression of See1 in maize
promoter of pit2, which is one of the U. maydis genes with the
cells, the effector was localized upon natural delivery (i.e., when
strongest in planta transcription (Skibbe et al., 2010; Doehlemann
it was secreted from infectious U. maydis hyphae). A C-terminal
et al., 2011). In leaf infections, the Ppit2-driven overexpression of
3xHA-tagged version of the effector was expressed under the
see1 RNA (verified by reverse transcription and quantitative PCR
control of the native promoter in the SG200Δsee1 deletion strain.
[RT-qPCR]) and protein (confirmed by immunoblot analysis) did
Immunoblot analysis after immunoprecipitation of See1-3xHA
not result in a phenotype significantly different from the wild type
from infected plant tissue verified the expression and stability of
Therefore, ectopic overexpression
the fusion protein ). For the immunoloc-
of see1 did not augment the virulence potential of U. maydis in
alization of See1-3xHA, plants infected with U. maydis strain
leaves. Interestingly, the see1-overexpressing strain caused an
SG200 served as negative controls. Two additional controls for
unexpected tassel phenotype, although tassel tumors caused
immunolabeling were employed: strain SG200 Psee1-GFP-3xHA
by wild-type U. maydis are largely restricted to the floral tis-
expresses cytoplasmic green fluorescent protein (GFP) driven by
sues, particularly the anthers. By contrast, Ppit2-driven see1
the see1 promoter, and strain SG200 Psee1-SPsee1-mCherry-
expression caused extensive tumor formation in the vegetative
3xHA expresses a secreted mCherry under the control of the
tassel base (the terminal node and internode) (Figure 5A). This
see1 promoter. Maize leaves were inoculated, and samples were
effect, which resulted in bizarre alterations of tassel architec-
harvested at 6 DPI for immunogold detection of the 3xHA tag. In
ture, was observed in ;38% of the infected plants (wild type,
the TEM images, no gold labeling was seen in plant tissue in-
8%) Additionally, ;20% of
fected with the parental strain SG200, indicating the absence
infected tassels including the spikelets became green 10 DPI
of nonspecific background labeling (Figure 6B, left panel). The
with the see1-overexpressing strain (
nonsecreted GFP-3xHA was detected exclusively inside fungal
hyphae at established biotrophic interfaces (Figure 6B, middle
Tissue infected by the see1-overexpressing strain was also
panel). The secreted mCherry-3xHA control showed labeling
used to quantify EdU-labeled cells in anthers as well as the
mainly in the biotrophic zone surrounding fungal cells as
vegetative tassel base. In anthers, overexpression of see1 did
well as inside fungal hyphae (Figure 6B, right panel). By
not cause any significant differences in EdU labeling compared
contrast, See1-3xHA was detected in the fungal hyphae, at the
Tumor Induction by Ustilago maydis
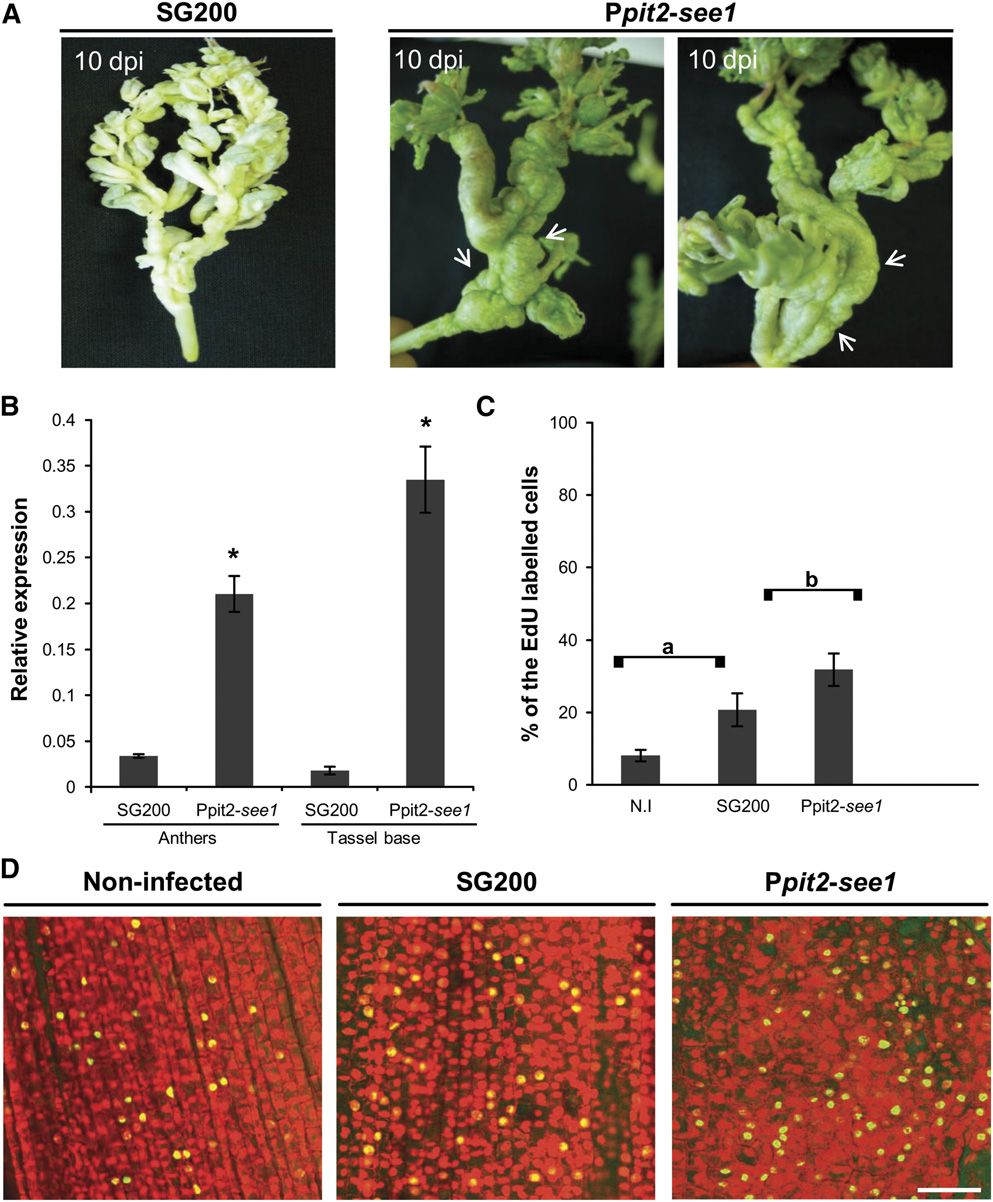
Figure 5. Overexpression of see1 Results in Tumor Proliferation in Vegetative Tassel Parts.
(A) Tassel base abnormality occurs much more frequently with constitutive overexpression of see1 in comparison with the wild-type strain SG200.
Tumor formation in the tassel base is indicated by the white arrows.
(B) Quantification of see1 gene expression in tassels infected with the overexpressing strain Ppit2-see1 in comparison with plants infected with the wild-type SG200 strain. Error bars show SE. *P # 0.001.
(C) Quantification of EdU-labeled tassel base cells in the in vivo DNA synthesis assay after infection with the See1-overexpressing strain Ppit2-see1 incomparison with either the wild-type SG200 infected or noninfected (N.I) tassels. There was a significant difference in the number of EdU-labeled nuclei inthe abnormal tassel base region as compared with the wild-type SG200 infected or noninfected tissue. Error bars show SE. Comparisons a and b, P # 0.05.
(D) Detection of in vivo DNA synthesis in the tassel base colonized by See1-overexpressing strain Ppit2-see1 in comparison with tissue colonized bywild-type strain SG200 and noninfected tissue. The total nuclei were visualized by PI staining (red), and the EdU-labeled cell nuclei were visualized byAF488 staining (green). Bar = 50 mm.
biotrophic interface, in plant cytoplasm, and prominently inside
found in the biotrophic interface. Only See1-3xHA was quan-
plant cell nuclei (Figure 6C; The dis-
titatively detected inside host cells, with ;20% of particles
tribution of gold particles was quantified for all constructs.
localizing to maize nuclei (Figure 6D). These results conclu-
While all the encoded proteins were found inside fungal cells,
sively demonstrate the translocation of See1 from biotrophic
only the two proteins with N-terminal secretion signals were
fungal hyphae into maize cytoplasm and nuclei.
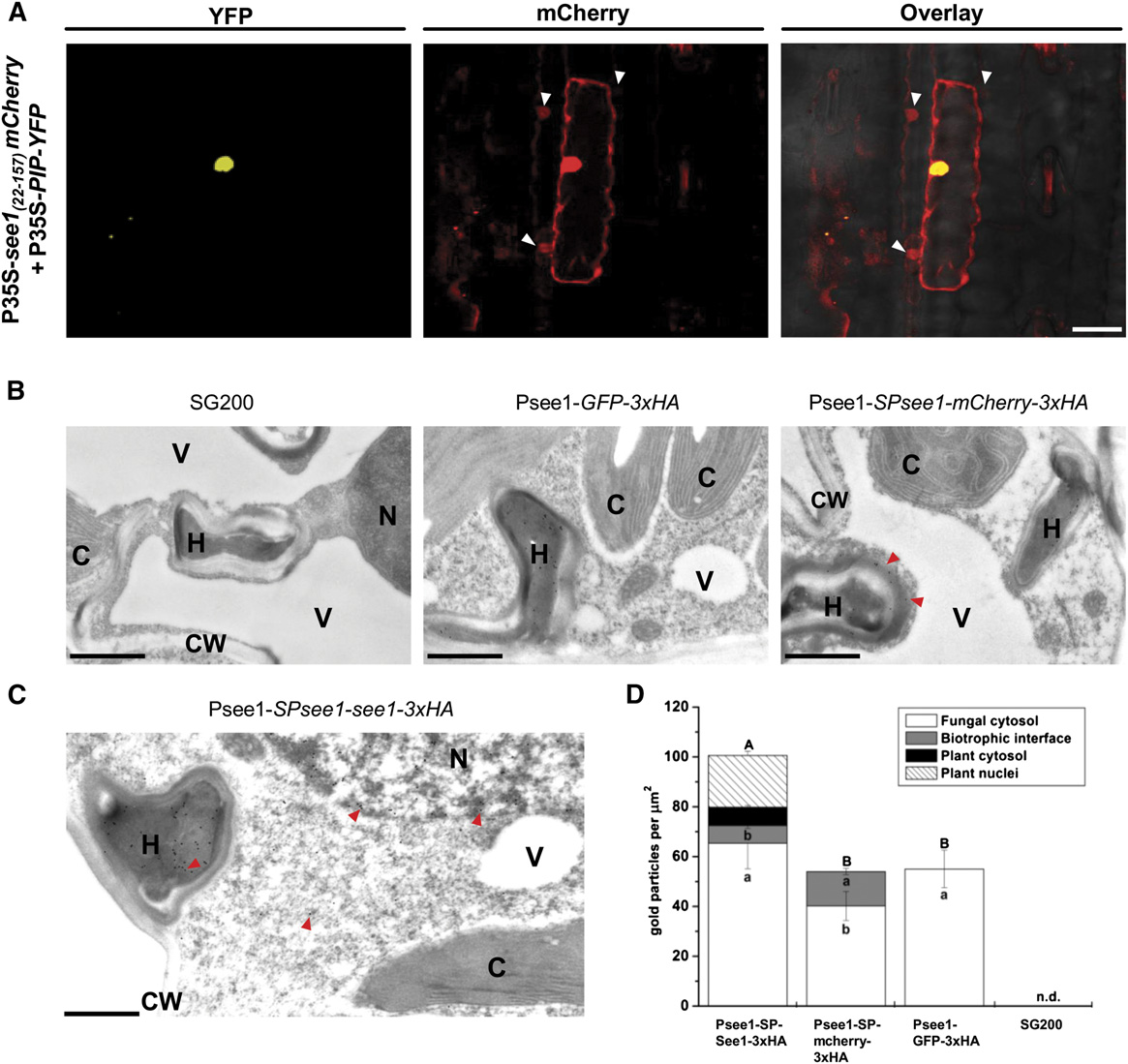
Figure 6. See1 Localizes to the Plant Cell Cytoplasm and Nucleus.
(A) Confocal microscopy of 35S-see1
-mCherry transiently expressed in maize epidermal cells. Left panel, transformation with the PIP-YFP control
results in fluorescence that is specifically localized to the nucleus. Right panel, See1-mCherry is localized to the cytoplasm and nucleus and istransferred to the adjacent neighboring cells, which are shown by the white arrowheads in the mCherry and overlay channels. Bar = 25 mm.
(B) Controls for the TEM micrographs showing immunogold labeling of See1-3xHA in leaves of U. maydis-infected maize. No gold particles were boundto wild-type infected tissue specimens (left panel). Gold labeling was restricted to fungal hyphae in GFP-3xHA samples, as GFP was not secreted by thefungus (middle panel). Gold particles bound to the secreted mCherry control could be found in hyphae and at the biotrophic interface (red arrowheads)but not inside the plant cells, despite proximity to hyphae. Psee1-SPsee1-mCherry-3xHA expression demonstrates that mCherry is secreted by thefungus but not taken up by the plant (right panel). Bars = 1 mm.
(C) Immunogold labeling of See1 could be found in hyphae (H), the cytosol, and nuclei (N), as shown by the red arrowheads, but not in chloroplasts (C),vacuoles (V), or the cell wall (CW) when the See1 effector was tagged with 3xHA in the strain Psee1-SPsee1-See1-3xHA. Bar = 1 mm.
(D) Graph depicts the spatial distribution of gold particles bound to See1-3xHA in different cell compartments of leaves from Psee1-See1-3x-HA alongwith the secretory (mCherry-3xHA), nonsecreted (GFP-3xHA), and SG200 wild-type controls. Means are shown with SE for the number of gold particlesper mm2 in the individual cell compartments of three independent transverse sections. Lowercase letters indicate significant differences (P < 0.05)between the individual cell compartments, whereas uppercase letters indicate significant differences between the total sum of labeling signal for allanalyzed cell compartments. Data were analyzed with the Kruskal-Wallis test followed by post-hoc comparison according to Conover (1999). n.d., notdetected, for all analyzed cell compartments.
Tumor Induction by Ustilago maydis
See1 Interacts with a Maize Homolog of SGT1
See1 Interferes with the Phosphorylation of SGT1
To identify proteins interacting with See1, we performed a yeast
A major question arising from the See1 interaction with SGT1
two-hybrid screen using a normalized cDNA library of U.
concerns how the effector interferes with SGT1 function at the
maydis-infected maize leaves and tassels. From 60 clones that
molecular level. Recent work showed that activation of SGT1
were isolated after plating on high-stringency selection medium,
signaling activity requires the phosphorylation of SGT1 by sali-
sequences corresponding to a maize homolog of SGT1 were
cylic acid-induced protein kinase (SIPK), a MAPK activated in
identified (Figures 7A and 7B). SGT1, a known regulator of cell
response to pathogen assault (Hoser et al., 2013). SIPK-mediated
cycle progression in yeast and an important factor in plant host
phosphorylation of SGT1 was concluded to trigger enhanced
and nonhost resistance, has three functional domains: the tetra-
nuclear compartmentalization of SGT1, thereby possibly activat-
tricopeptide repeat (TPR) domain, the Chord SGT1 (CS) domain,
ing defense-related signaling via the modulation of transcription.
and an SGT1-specific (SGS) domain (Figure 7A) (Kitagawa et al.,
Based on these observations, we tested whether See1 interferes
1999; Peart et al., 2002). There are also two variable protein re-
with the phosphorylation of SGT1. To facilitate the analysis of the
gions that are species-specific (Figure 7A). To test whether the
in planta phosphorylation of SGT1 in the presence of See1, we
identified maize protein exhibits SGT1 functions, temperature-
used the well-established Agrobacterium tumefaciens expression
sensitive mutants of S. cerevisiae, YKK57 (sgt1-5) and YKK65
system in N. benthamiana. N. benthamiana SGT1, which shares
(sgt1-3) (Kitagawa et al., 1999), were used for complementation
65% identity to maize SGT1 ), was pre-
experiments. At 37°C, both mutants are restricted in cell cycle
viously shown to be activated by Nt-SIPK, which, in turn, shares
phases, arrested at G1 and G2, respectively. Expression of full-
84% to 86% identity to a set of five putative MAPK proteins of
length maize SGT1 under the control of the GAL4 yeast promoter
maize ). Agrobacterium strains carrying
complemented the growth defect of S. cerevisiae strain YKK57
P35S-Zm-SGT1-StrepII, a gene encoding constitutively active
(sgt1-5), indicating the functionality of the identified maize ho-
Nt-MEK2 (Nt-MEK2DD) under the control of a dexamethasone
molog. Expression of maize SGT1 in S. cerevisiae strain YKK65
(DEX)-inducible promoter, P35S-SIPK, and Dex-see1-HA or pTA7001
(sgt1-3), which is defective at G2, showed normal growth at
empty constructs were coinfiltrated into N. benthamiana leaves
permissive temperature and partial complementation at 37°C
and maintained for 2 d to ensure expression of the constitutive
promoter constructs. Subsequently, Nt-MEK2DD and See1 ex-
To verify the See1-SGT1 interaction in planta, both proteins
pression were induced by DEX treatment. Leaf samples were
were transiently expressed in Nicotiana benthamiana. As expression
collected 5 h after the induction, and SGT1 was affinity-purified
controls, P35S-See1-Myc and P35S-SGT1-HA were separately
via its Strep tag II. Expression of all heterologous proteins has
expressed in N. benthamiana leaves (Figure 7C). Using anti-HA
been verified by immune detection We
matrix, See1-Myc was coimmunoprecipitated by the HA-tagged
detected two sites of phosphorylation in Zm-SGT1. Constitutive
SGT1 but not in the absence of SGT1 (Figure 7C), confirming
phosphorylation that was independent from See1 as well as
the See1-SGT1 interaction in planta. To localize the See1-
from SIPK was detected for residue Thr-262
SGT1 interaction in plant cells, bimolecular fluorescence
and This residue is situated in
complementation (BiFC) was employed, using an enhanced
the second variable region of Zm-SGT1 and only conserved in
split-YFP system (Hemetsberger et al., 2012). An mCherry tag
maize and sorghum (Sorghum bicolor)
was fused to the C terminus of the N-terminal part of YFP
Surprisingly, upon SIPK activation and in the absence of See1,
(pSPYNE_N). Similarly, a cyan fluorescent protein (CFP) tag
there was a high abundance of a Zm-SGT1 phosphopeptide
was added to the C-terminal part of YFP (pSPYCE_C). Via
with phosphorylation at Thr-150 (Figures 8A and 8B) in all three
ballistic transformation of maize epidermal cells, both con-
independently performed biological replicates. This phosphor-
structs were transiently expressed under the control of the 35S
ylated position is situated in the variable region of Zm-SGT1,
promoter. Cells expressing both pSPYCE_C and pSPYNE_N
represents a putative MAPK target site, and is conserved in
fused to SGT1 and See1, respectively, were designated as
monocots including maize, rice (Oryza sativa), and sorghum
). Strikingly, in See1 coinfiltrated samples,
CFP-CYFP-HA. The cells exhibited cytoplasmic and nuclear
no phosphorylation at Thr-150 could be detected, reflecting the
fluorescence signals for both mCherry and CFP, indicating
interference of See1 with SIPK1-induced phosphorylation of SGT1
expression of the fusion proteins. Expression of pSPYNE_N-
and ). As a negative
mCherry with pSPYCE_SGT1 did not result in any detectable
control, we coexpressed an inactive form of Nt-MEK2 (Nt-MEK2KR)
YFP signal, demonstrating that no unspecific protein di-
with SGT1. In this case, SIPK was not included because its tran-
merization occurred (Figure 7D). Similarly, no YFP fluorescence
sient expression results in slightly increased SIPK activity and the
was detected when pSPYCE-CFP was coexpressed with
associated activation of defense responses (Zhang and Liu, 2001).
pSPYNE-mCherry fused to see1 (pSPYNE_see1) (Figure 7D). By
Under these conditions, coexpression of Nt-MEK2KR and SGT1 did
contrast, cells that coexpressed pSPYNE_see1 and pSPYCE_SGT1
not result in SGT1 phosphorylation at Thr-150. Therefore, we
showed a complementation of YFP fluorescence (Figure 7D),
conclude that See1 interferes with the Zm-SGT1 phosphorylation at
indicating an interaction of See1 and SGT1 in the cytoplasm
Thr-150, thereby delaying or preventing its activity.
and nucleus of maize cells. Altogether, we demonstrate that
To corroborate the result that SIPK could phosphorylate Zm-
See1 interacts with SGT1 in the cytoplasm and nucleus of
SGT1, purified recombinant proteins Zm-SGT1, SIPK, and MEK2DD
maize cells.
were coincubated in the presence of radioactive ATP in vitro. As
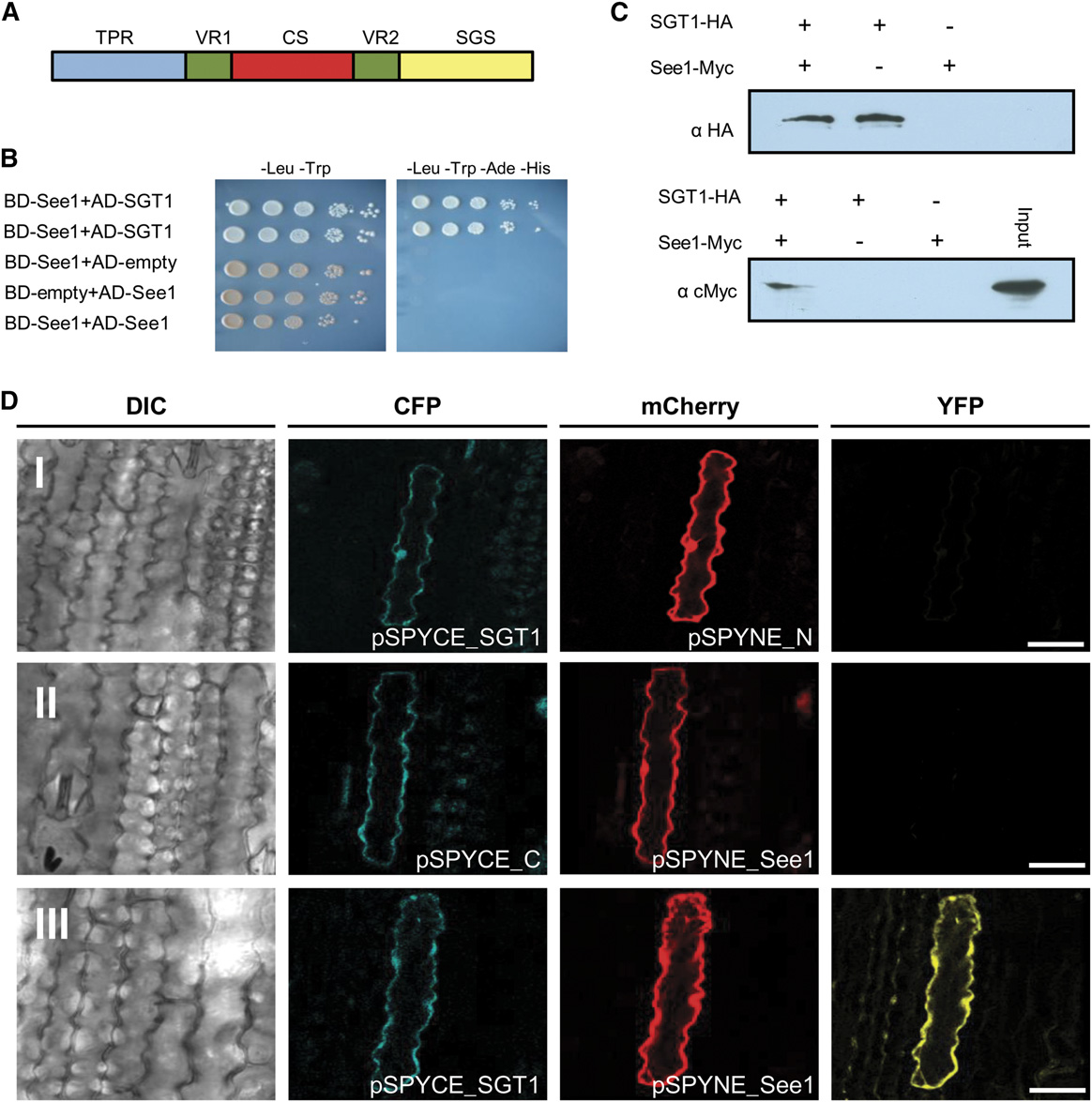
Figure 7. See1 Interacts with the Cell Cycle and the Immune Response Modulator SGT1.
(A) Domain structure of maize SGT1 depicting three important domains: TPR, CS, and SGS. The two variable regions (VR1 and VR2) in the protein arespecies-specific.
(B) Yeast two-hybrid experiment to test for the interaction of See1 and maize SGT1. The drop assay was done by serial dilutions (see Methods), andstrains were tested on low- and high-stringency plates to check for the specificity of the interaction. Results were documented after 4 d.
(C) Coimmunoprecipitation shows the interaction of See1 and SGT1 fusion proteins isolated from transiently expressing N. benthamiana cells. SGT1was tag purified, and See1 was pulled down. In the absence of SGT1, no See1 signal was detected.
(D) In vivo interaction of See1 with SGT1. Confocal images show maize epidermal cells expressing BiFC constructs. Row I shows a plant cellcoexpressing pSPYCE-SGT1 and pSPYNE-mCherry. Blue and red channels show cytoplasmic colocalization of the respective signals. No comple-mentation of fluorescence is observed in the YFP channel. Row II shows the coexpression of pSPYCE-CFP and pSPYNE-See1. Blue and red channelsshow cytoplasmic colocalization of the respective signals. No complementation of fluorescence is observed in the YFP channel. Row III shows a cellcoexpressing pSPYCE-SGT1 and pSPYNE-See1. Both signals colocalize in the nucleus and cytoplasm. The YFP channel exhibits YFP fluorescencereflecting the direct interaction of See1 and SGT1. DIC, differential interference contrast. Bars = 25 mm.
shown in the SIPK activated by
a positive control, a nonspecific substrate, myelin basic protein
MEK2DD was able to phosphorylate Zm-SGT1 (lane 1). In the
lane 3), was used; this protein was in-
negative control, which did not contain SIPK, we did not observe
tensively phosphorylated in our assay. Collectively, from these
any phosphorylation of Zm-SGT1, and the signal detected cor-
results, we conclude that SIPK from tobacco (Nicotiana tabacum)
responded to MEK2DD (, lane 2). As
can phosphorylate Zm-SGT1 in planta or in vitro and, therefore,
Tumor Induction by Ustilago maydis
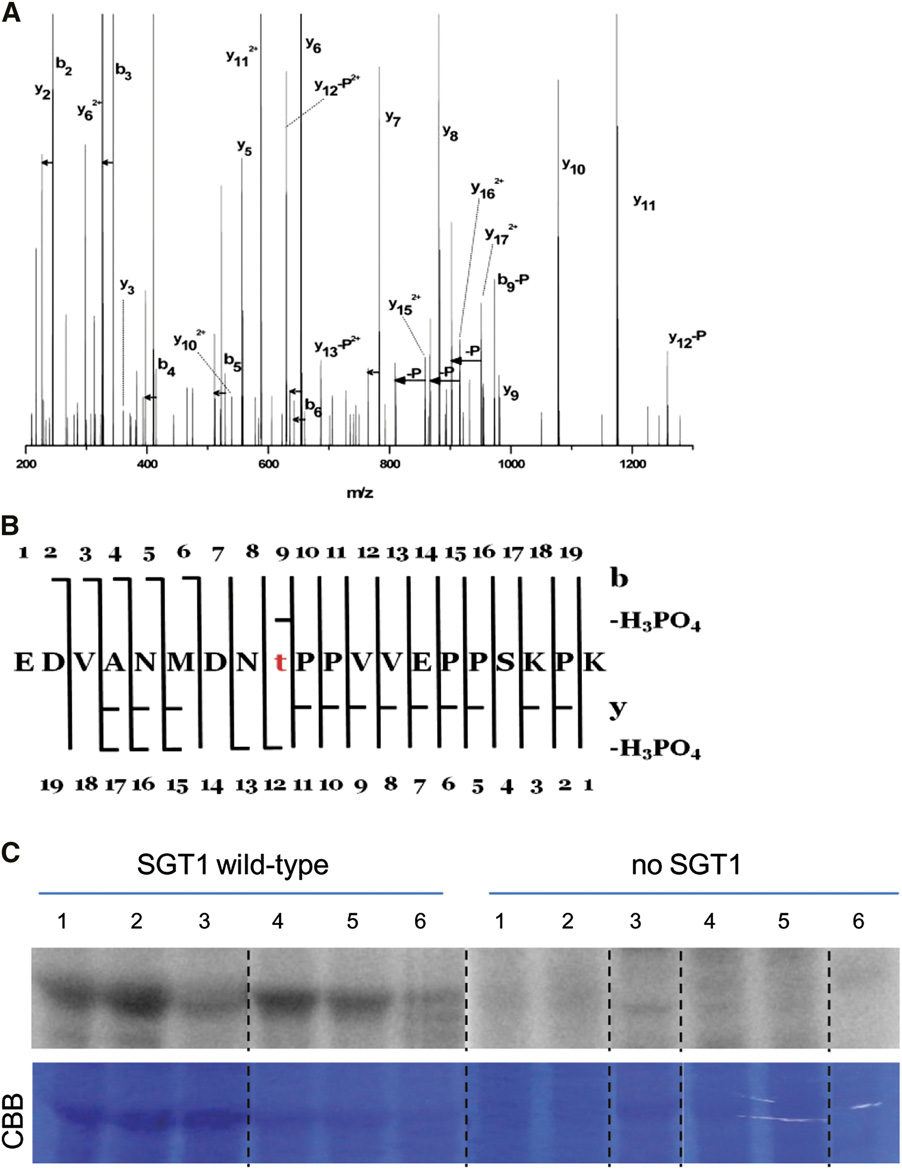
Figure 8. In Planta Phosphorylation of Maize SGT1.
(A) Fragmentation spectrum assigned to the phosphorylated form of the peptide EDVANMDNTPPVVEPPSKPK (Mascot score 126). Loss of H PO is
denoted by –P, loss of water is marked by short horizontal arrows, whereas a longer arrow symbolizes pairs of detected signals corresponding to y and
y -H PO . The majority of signals of the tandem mass spectrometry spectra are assigned to the above species. The presence of several y
and b -H PO ions accompanied by y , y , and y
pinpoints threonine at position 9 as the unequivocal phosphorylation site within the peptide.
(B) Peptide sequence with assigned y, b, y-H O, b-H O, y-H PO , and b-H PO ions present.
(C) Recombinant maize SGT1 produced in E. coli was incubated in the buffer containing [g-32P]ATP, and total proteins were extracted from maize seedlingsor tassels infected by various U. maydis strains. The samples were fractionated by SDS-PAGE and analyzed with a phosphor imager. Columns 1 to 3,extracts from seedling leaves 6 DPI with U. maydis wild-type SG200 (1), SG200Δsee1 (2), or mock-inoculated (3). Columns 4 to 6, extracts from tassel base9 DPI with U. maydis wild-type SG200 (4), U. maydis-overexpressing Ppit2:see1 (single-copy integration; 5), or U. maydis-overexpressing Ppit2:see1(multiple-copy integration; 6). Representative data of four independent biological replicates are shown. CBB, Coomassie Brilliant Blue.
could be used for transient assays to study the See1-SGT1
while a phosphomimic variant of At-SGT1b exhibited enhanced
nuclear localization (Hoser et al., 2013). To test whether the
Phosphorylation can alter protein subcellular localization and
phosphorylation of Zm-SGT1 altered its subcellular localization,
stability (Cohen, 2000). A truncated Arabidopsis thaliana SGT1b
constructs encoding YFP fused C-terminally to the wild-type
lacking a MAPK phosphorylation site was detected only in the
Zm-SGT1 or to its phosphovariants were transiently expressed
nucleus-depleted fraction in leaf extracts (Noël et al., 2007),
in N. benthamiana epidermal cells via particle bombardment.
Wild-type Zm-SGT1 localized to the nucleus in ;33% of the
constructs), tassel bases showed quantitative reduction of SGT1
transformed cells, and its relative nuclear fraction in all cells
phosphorylation (Figure 8C). When using Zm-SGT1 protein with
tested was 39.4% Upon bombard-
the T150A (SGT1AP) mutation in the same assay, residual phos-
ment with constructs encoding Zm-SGT1 fused to a strong
phorylation was observed mainly in the tassel tissue
nuclear localization signal derived from SV40 large T antigen or
. This signal did not correlate with See1 expression and,
protein serine kinase protein-derived nuclear export signal, the
therefore, most likely represents the previously observed consti-
nuclear fraction was changed to 94.3% and 15% of the trans-
tutive phosphorylation of residue Thr-262 (
formed cells, respectively (Wen et al., 1995). Control-like subcellular
and Together, these findings demon-
distribution could be restored by mutation of either the nuclear lo-
strate that Thr-150 phosphorylation of maize SGT1 is triggered in
calization signal or the nuclear export signal
response to U. maydis infection and that this is modulated by the
, demonstrating the suitability of the bombardment assay to
See1 effector.
assess protein localization. The phosphonull (Zm-SGT1AP) variantlocalized to the nucleus in around 20% of the transformed cells,
and the nuclear fraction was 33.5% . Bycontrast, the phosphomimic variant of Zm-SGT1 (Zm-SGT1DP)
The secreted effector protein See1 is an organ-specific U. maydis
was observed in the nucleus of ;40% of transformed cells, with
virulence factor that promotes tumor formation in maize vegeta-
an average nuclear fraction of 46.2%. These data suggest that
tive tissues. The expression profile of see1 shows strong organ
Zm-SGT1 phosphorylation status affects nuclear import or export.
specificity, supporting the specific requirement for this effector in
Because the differences are subtle compared with the wild type, we
supporting tumors in maize leaves but not in floral tissues. See1 is
conclude that the disturbance in localization of the Zm-SGT1 is not
translocated from biotrophic hyphae to the maize cell cytoplasm
the primary virulence function of See1.
and nucleus, where it interacts with maize SGT1 and interfereswith its MAPK-induced phosphorylation. In leaf zones with post-mitotic differentiated cells, U. maydis requires See1 to reactivate
The See1-SGT1 Interaction Has Functional Relevance
host cell division, a prerequisite for tumor formation. By contrast,
To further investigate the role of See1 in targeting SGT1, we
anther tumor induction does not require de novo activation of
checked if the identified phosphorylation at Thr-150 in Zm-SGT1
plant cell proliferation, because in this highly proliferating tissue,
is required for SGT1 function. To address this point, we first
the tumors result from redirecting host cell division and cell ex-
performed a yeast complementation assay using sgt1 cell cycle
pansion into a tumor pathway (Gao et al., 2013).
mutants to test for the complementation ability of phosphomi-
During maize leaf development, most cell divisions occur in
mic and phosphonull mutants of Zm-SGT1 at this position. This
a narrow zone at the base of the blade adjacent to the ligule,
assay did not show any differences in the complementation
with only sporadic divisions in the differentiating leaves. Tumors
ability compared with the wild-type SGT1
formed in U. maydis infections result from the profuse and rapid
This suggests that the phosphorylation site we identified in
cell division in the subepidermal leaf cells. In uninfected plants,
planta is not required for the cell cycle-related SGT1 function in
no such activity was visible in the corresponding leaf area (which
yeast. Because the relevant phosphorylation site is specific to
is neither part of the apical meristem nor the basal region of the
monocot SGT1 homologs and not present in yeast, one could
leaf) (Li et al., 2010). Confocal microscopy of leaves infected by
not expect a function of this residue in yeast.
SG200Δsee1 showed that it successfully penetrated the host
To elucidate the relevance of SGT1 phosphorylation in tumor
tissue and established itself in the initial stages of colonization.
formation, an alternative approach was employed. Specifically,
During the later stages, when the fungi reached the mesophyll
this aimed to identify the phosphorylation status of SGT1 upon
cell layer, the mutant displayed defects in passing from cell to
U. maydis infection and to further test the phosphorylation state
cell, with entrapment of the fungal hyphae in the mesophyll cells
in overexpression strains as compared with the U. maydis wild-
or the adjacent vascular cells This stage
type SG200. To this end, maize leaves were infected with the
of infection at 4 DPI coincides with the normal appearance of
wild-type U. maydis strain SG200 and SG200Dsee1. In addition,
heavy EdU labeling, indicative of reinitiation of the cell cycle in
tassels were infected with strains overexpressing See1 (which
leaves infected with the wild-type fungi, as monitored by DNA
caused the abnormal tassel base phenotype) to collect the vege-
precursor incorporation (Gratzner, 1982; Salic and Mitchison,
tative tassel base tumors. An in vivo detection of SGT1 phos-
2008). EdU labeling of seedling tissue colonized by U. maydis
phorylation was performed by preparing extracts from infected
showed that several division events had occurred in contiguous
tissues, which served as a kinase source in the assay. Supple-
cells by 4 to 5 DPI, indicating that reactivation of host cell division
mental recombinant Zm-SGT1 in maize leaf or tassel base extracts
occurs after initial fungal establishment and is followed by sus-
was monitored for its phosphorylation state by autoradiography. It
tained proliferation of maize leaf cells.
is clear that Zm-SGT1 is phosphorylated after incubation with the
During the initial 2 d of anther colonization, U. maydis is present
extracts from U. maydis-infected leaves as compared with the
on the epidermis (Gao et al., 2013). At later time points, the fungus
uninfected control (Figure 8C). Moreover, incubation with the ex-
is subepidermal and alters cell fate specification events, ongoing
tracts from leaves infected with SG200Dsee1 showed increased
cell division patterns, and cell expansion depending upon de-
phosphorylation of SGT1 compared with the extracts from wild-
velopmental stage and cell type. The fungus mainly induces
type SG200-infected tissue. With U. maydis overexpression of
ectopic periclinal divisions in anther somatic cells, generating
See1 (by single and multiple integration of overexpression
an extra cell layer resulting in disrupted anther lobe architecture.
Tumor Induction by Ustilago maydis
Frequent anticlinal and periclinal divisions are also observed in
subsequent cell-to-cell movement may reflect a general feature of
the middle layers of infected anther, which otherwise undergo
effectors taken up by plant cells. These proteins could act to
only a few anticlinal divisions after their birth, prior to programmed
stimulate the surrounding cells not yet in contact with fungal hy-
cell death. Hence, in floral tissues, U. maydis reprograms cell fate
phae to promote fungal proliferation. For See1, the independent
but does not act as an oncogenic agent (Gao et al., 2013).
approach of immunolabeling clearly confirmed that the effector is
Constitutive overexpression of see1 led to tumor formation in the
translocated to the plant cell cytoplasm and nucleus. Movement
vegetative parts of the tassel base, which in wild-type infections
of See1 to the neighboring cells was not observed in the TEM
were not tumorous under the tested conditions. This suggests
immunogold assay. This discrepancy might result from different
that See1 specifically acts in vegetative tissues. The phenomenon
expression levels in the two approaches; in the immunolabeling,
might be of significance to the fungus in nature, where floral tu-
See1 was expressed by fungal hyphae from its native promoter,
mors are more frequent in occurrence. See1 and other leaf-
but plant-derived expression was driven by the 35S promoter.
specific effectors might be of importance for host adaptation and
Nevertheless, based on our data, we cannot exclude that the cell-
the evolution of U. maydis, as they promote the formation of tu-
to-cell movement observed in the transient assay is an experi-
mors in vegetative parts, an important factor in colonizing pe-
mental artifact of the exogenous expression of the effector rather
rennial grasses or exploiting seedlings. Infection of seedlings and
than a feature of normal infections. On the other hand, the phe-
immature plants allows the fungus to complete its short, 2-week
nomenon of cell-to-cell movement of translocated effectors has
life cycle multiple times during plant vegetative growth, because
also been described for Magnaporthe oryzae effectors in rice cells
this infection style is independent from the development of plant
(Khang et al., 2010). This movement between cells was proposed
to occur via the plasmodesmata, which are coopted by hyphae for
Where does the See1 effector act in leaf cells? Transient ex-
cell-to-cell movement (Kankanala et al., 2007; Djamei et al., 2011).
pression assays showed that See1 localizes to the cytoplasm
See1 interacts with maize SGT1 in a yeast two-hybrid screen.
and nucleus of maize cells. Interestingly, the effector protein
Maize SGT1 is present in both the cytoplasm and nucleus of
also moved to cells neighboring a transformed cell. The specific
plant cells; hence, localization is consistent with the dual locali-
translocation of See1-mCherry between neighboring maize cells
zation of See1. The interaction of See1 and SGT1 was confirmed
suggests that there may be a second route for fungal effectors
independently by in planta coimmunoprecipitation. In addition,
to enter maize cells. The U. maydis effectors Cmu1 and Tin2
BiFC data showed that this interaction takes place in both the
are both translocated from biotrophic hyphae into the host
cytoplasm and nucleus of maize cells. This evidence is congruent
cell (Djamei et al., 2011; Tanaka et al., 2014). We speculate that
with the electron microscopy analyses that show the same
Figure 9. Tentative Model of the Role of the See1-SGT1 Interaction during U. maydis Tumor Formation.
The SGT1 protein is known to occur in the cytoplasmic and nuclear pools (Hoser et al., 2013). In U. maydis wild-type infections, activated unidentifiedmaize kinase (UMK) triggers the phosphorylation of SGT1 at a monocot-specific target site. The See1 effector binds to SGT1, interferes with itsphosphorylation status, and thereby disturbs the subcellular distribution (i.e., transport into the nucleus). This misbalancing of SGT1 phosphorylationand distribution contributes to the induction of cell cycle genes, leading to the induction of tumorous division.
localization pattern of the See1 effector. Mass spectrometry
for symptom development. The precise steps following the in-
analysis showed that the interaction with See1 results in an in-
terference of See1 with the posttranslational modification of
hibition of SGT1 MAPK-triggered phosphorylation at Thr-150.
SGT1, resulting in the reactivation of maize DNA synthesis and
Why would See1 interfere with the phosphorylation of SGT1
ultimately in tumor formation, remain to be elucidated biochemi-
during leaf infection? Hijacking of SGT1 may contribute to the
cally. Therefore, it will be of prime interest to work out the detailed
deactivation of immune responses. The literature provides a large
molecular mechanism of the See1-SGT1 interaction and to dissect
body of data on SGT1. From all the available evidence concerning
the downstream signaling network (i.e., to identify and characterize
plant SGT1, this protein is widely seen to be active in vegetative
proteins that interact with and/or are affected by SGT1 and to
leaf tissues (Noël et al., 2007). In Arabidopsis, the SGT1b isoform
pinpoint residues in See1 required to interfere with SGT1).
was found to be required for the SCF-mediated auxin response in
Transcriptome analysis of U. maydis wild-type and SG200Δsee1-
seedling roots (Gray et al., 2003). Arabidopsis SGT1 has a regu-
infected maize leaves at 6 DPI showed that genes involved in
latory role in early R gene-mediated plant defenses (Austin et al.,
DNA binding, replication (including Skp1), as well as repair mech-
2002) and was shown to be involved in forming a cochaperone
anisms characterize normal infections but are not induced in the
complex with HSP90; this complex functions in sensing immune
mutant interaction. We hypothesize that this reflects the reduced
responses of the host receptor proteins (Shirasu, 2009). Among
activation of cell division and, consequently, limited tumor expan-
the three domains of SGT1, namely the TPR, CS, and SGS do-
sion in the mutant-infected leaves. Recent work by Bao et al. (2013)
mains, the CS domain resembles the a-crystalline domain of the
found an unexpected link between cell cycle progression and plant
cochaperone HSP20 (Dubacq et al., 2002; Garcia-Ranea et al.,
immunity, suggesting that cell cycle misregulation impacts the ex-
2002). The other components of the immune regulatory cocha-
pression of R genes. Also, DNA repair proteins have been shown to
perone complex, RAR1 and HSP90, have been shown to interact
be directly involved in the regulation of gene expression during
with the CS domain of SGT1 (Azevedo et al., 2002; Takahashi
plant defense responses (Song et al., 2011). The DNA damage
et al., 2003).
response is an intrinsic component of the plant immune response
The host SGT1/RAR1/HSP90 complex is a target of several
and, in turn, enhances salicylic acid-mediated defense gene ex-
bacterial effector proteins. The Pseudomonas syringae effector
pression (Yan et al., 2013). Among the Ustilaginales, which cause
AvrB interacts weakly with Arabidopsis SGT1b (Cui et al., 2010).
characteristic floral symptoms in the immature, proliferative host
Another P. syringae effector, AvrPtoB, showed a genetic in-
floral organs, U. maydis is the only species that causes local tumors
teraction with SGT1 and RAR1, requiring these cochaperones to
in vegetative tissue. Seedling-specific effectors such as See1 pro-
suppress plant immunity (Hann and Rathjen, 2007). Additionally,
mote the generation of a mitotically active sink tissue within vege-
the P. syringae effector HopI1 interacts with HSP70 (Jelenska
tative organs. By regulating SGT1, See1 may not only shut down
et al., 2010), which is an active component initiating signaling by
defense signaling but also activate the host cell cycle, a prerequisite
interaction with the SGT1/RAR1 complex. Recently, effector
for tumor development. Hence, a combination of immune sup-
proteins from Salmonella enterica and Xanthomonas campestris
pression and nutrient rechanneling, particularly facilitating the ac-
have been shown to interact with SGT1 (Bhavsar et al., 2013;
quisition of sucrose, could trigger uncontrolled cell proliferation,
Kim et al., 2014). Consistent with the previous findings, these
ultimately resulting in plant tumors. Based on the results obtained in
effectors bind to SGT1 at the CS domain, confirming the impor-
this study, our hypothesis on how the See1-SGT1 interaction could
tance of this domain for SGT1 regulation during immune de-
affect the formation of leaf tumors is shown in a tentative model
fenses. SGT1-mediated pathways may vary by plant species and
(Figure 9). Detailed understanding of such processes should shed
are also specific to a particular pathogen (Wang et al., 2010).
light on colonization biology in various biotrophic plant pathogens
SGT1 is known to be used by some fungal pathogens in pro-
and on intrinsic host mechanisms that operate to prevent cell
moting disease symptoms. Like X. campestris, the necrotrophic
proliferation in differentiated organs.
fungus Botrytis cinerea uses SGT1 to initiate the hypersensitiveresponse-mediated cell death pathway for the necrotrophic life-style (El Oirdi and Bouarab, 2007). Fusarium culmorum is known
to require SGT1b to cause full disease symptoms in buds andflowers of Arabidopsis (Cuzick et al., 2009), the only prior report
Growth Conditions and Virulence Assays
that described SGT1 in floral tissues. These observations suggest
Maize (Zea mays cv Early Golden Bantam [Olds Seeds] and cv Gaspe Flint
that searching for a tissue-specific role of SGT1 protein in Ara-
[maintained by self-pollination]) plants were grown in a temperature-
bidopsis flowers may be fruitful.
controlled greenhouse (14-h/10-h light/dark cycle, 28/20°C). Gaspe Flint
We propose that SGT1 represents a conserved hub targeted
plants were mainly used for tassel infection and for the overexpression of
by several effectors from bacterial as well as fungal pathogens,
See1, as they have an early floral switch (15 d) and are suitable for early
utilizing it according to the need of the pathogen lifestyle. This
meristematic tassel infections. Both cultivars of maize exhibit similar
supports the early study of evolutionarily different effectors tar-
infection symptoms. Both varieties were grown in T-type soil (FrühstorferPikiererde). Ustilago maydis strains were grown in YEPS (0.4% yeast
geting a common host defense protein (Song et al., 2009) and is
extract, 0.4% peptone, and 2% sucrose) at 28°C with shaking at 200 rpm
also consistent with the model of evolutionarily different virulence
of 0.6 to 0.8. Cells were centrifuged at 900g for 5 min, re-
effectors targeting conserved hubs in a plant immune system
suspended in water to OD
of 1.0, and injected into stems of 7-d-old
network (Mukhtar et al., 2011). Work to date demonstrated that
maize seedlings with a syringe, as described previously (Kämper et al.,
SGT1 is involved in resistance during biotrophic interactions. The
2006). All infection assays were performed in three independent infection
results reported here indicate that U. maydis alters SGT1 function
trials with multiple seedlings. Disease symptoms were scored at 12 DPI
Tumor Induction by Ustilago maydis
using a previously developed scoring scheme (Kämper et al., 2006).
and quick penetration of the fixer and allows better preservation of mitotic
Tassel infections were done after 15 d of seed sowing in Gaspe Flint and
stages (Kotogány et al., 2010).
after 4 weeks in Early Golden Bantam, as described previously (Walbotand Skibbe, 2010). Disease symptoms in the tassels were scored 14 DPI
Yeast Transformation and Two-Hybrid Interaction Assay
following the criteria described in a previous study (Schilling et al., 2014).
Nicotiana benthamiana plants (BN3) were grown in a phytochamber
To identify host interactors of See1, See1 lacking the signal peptide was
(Vötsch) under controlled environmental conditions (21°C, 16 h of light, 8 h
expressed from pGBKT7-See1
and screened against a maize cDNA
of dark) as described previously (Talarczyk et al., 2002).
library in pGADT7. Preliminary testing indicated that See1 was nontoxic toyeast (Saccharomyces cerevisiae) and did not autoactivate. The yeasttwo-hybrid library analyses were done using a normalized cDNA library of
RNA Extraction and RT-qPCR
infected maize tissue containing seedling and tassel samples. The strain
Expression of the candidate gene was analyzed by RT-qPCR. Total RNA
AH109 (Clontech) was used for all yeast assays unless otherwise men-
was isolated from infected samples (leaf, tassel base, or anthers). The
tioned. Yeast transformation was done as described in the DUAL
infected plant material was collected at successive time points from 2 to
membrane starter kit manual (Dualsystems Biotech). The yeast two-hybrid
14 DPI. To confirm the high expression of see1 at 9 DPI, seedlings, anthers,
screen was performed following the instructions of the Matchmaker
and tassel base were collected from plants infected with U. maydis strain
yeast two-hybrid manual (Clontech) using 1 mg of bait DNA (pGBKT7-
Ppit2-see1 and the SG200 control. The samples were taken in three in-
See1) and 0.5 mg of library DNA. All resulting yeast clones were tested by
dependently conducted experiments. RNA was isolated using Trizol re-
immunodetection for expression of the respective proteins. To perform
agent (Invitrogen) and purified using an RNeasy kit (Qiagen). For cDNA
a yeast dilution assay, 3 mL of selective medium (SD-Leu-Trp) was in-
synthesis, the SuperScript III first-strand synthesis SuperMix kit (Invitrogen)
oculated with a single colony of the respective yeast strain and incubated
was used to reverse transcribe 1 mg of total RNA with oligo(dT) primer. The
overnight at 28°C. OD
was adjusted to 0.2, and the cells were grown to
RT-qPCR analysis was performed using an iCycler machine (Bio-Rad) in
of 0.6 to 0.8. Next, 1 mL of yeast culture was centrifuged for 10
combination with iQ SYBR Green Supermix (Bio-Rad). Cycling conditions
min at 3500g, and the pellet was washed twice with 1 mL of sterile water
were 2 min at 95°C followed by 45 cycles of 30 s at 95°C, 30 s at 61°C,
and finally resuspended in 500 mL of sterile water. OD
and 30 s at 72°C. Gene expression levels were calculated relative to
1.0 with sterile water, and 5 mL of this suspension, as well as 1:10, 1:100,
the peptidylprolyl isomerase gene (ppi ) of U. maydis for quantifying
and 1:1000 dilutions, were applied on SD-Leu-Trp plates (low stringency)
see1 expression (van der Linde et al., 2012). Error bars in all figures that
as a growth control and SD-Leu-Trp-Ade-His plates (high stringency) to
show RT-qPCR data represent the SD that was calculated from the
test for protein-protein interaction. Growth was scored after 4 to 5 d of
original cycle threshold values of three independent biological repli-
incubation at 28°C.
cates. Primer sequences used for RT-qPCR are listed in
For the yeast complementation assay of the yeast sgt1 mutants, yeast
strains YKK 57 (sgt1-5) and YKK65 (sgt1-3) were transformed with con-structs to express maize SGT1 and the phosphovariants of SGT1 (cloned
EdU-Based DNA Synthesis and Cell Proliferation Assay
into pGREG536 vector under the control of the Gal1 promoter). Thetransformation was done as described previously, and the transformants
Published protocols were used for U. maydis seedling infections (Kämper
were selected on SC-Ura-2% (w/v) glucose plates. The strains were then
et al., 2006) and tassel infections as described (Walbot and Skibbe, 2010).
shifted to SC-Ura-2% (w/v) galactose and incubated for 4 d to test their
At 4 DPI, the third infected leaf where the first infection symptoms ap-
ability to complement the temperature-sensitive sgt1-5 and sgt1-3 growth
peared was used for the EdU assay and incubated for 5 h with 10 mM EdU
defects. Also, the transformants selected were serially diluted 5-fold
(Invitrogen) in small chambers designed for labeling physiologically active
for a drop assay and incubated at 25 and 37°C for 4 d to check for
leaves. For the tassels at 3 DPI, the immature mitotically active tassel was
bathed with 1 mL of 20 mM EdU after delivery with a 26-gauge hypodermicneedle through the whorl of leaves surrounding the inflorescence apex
;17 d after germination in the Gaspe Flint maize variety. EdU seeps intospikelets through the small air spaces between the external organs of the
Confocal images were taken on a TCS-SP5 confocal microscope (Leica)
spikelet and florets and reaches anthers over the labeling time of 5 h. After
as described previously (Doehlemann et al., 2009). Details of the AF488
the labeling procedure, the area in seedlings below the infection holes was
WGA and PI microscopy are given in . Fluores-
detached and fixed in 100% (v/v) ethanol. For the tassel tissue, around
cence of AF488 alone or coupled to EdU was elicited at 488 nm and
150 anthers from different parts of the tassel were dissected to ensure
detected at 495 to 540 nm. PI was excited at 561 nm and detected at 570
random sampling with equal probability of labeled anthers and fixed in
to 640 nm. YFP was excited at 514 nm and detected at 520 to 540 nm,
100% (v/v) ethanol 5 h after labeling, as done for seedlings. The EdU
mCherry fluorescence was excited at 561 nm and detected at 590 to 630
staining procedure was done as described previously (Kelliher and
nm, and cell wall autofluorescence used excitation of 405 nm and de-
Walbot, 2011). The samples were washed once with fresh 100% (v/v)
tection at 435 to 480 nm.
ethanol followed by two washes in PBS, pH 7.4, plus 2% (w/v) BSA, then
Preparation of samples for TEM and immunogold labeling was per-
the samples were transferred to permeabilization solution (PBS +1% [v/v]
formed as described previously (Heyneke et al., 2013). Small samples
Triton X-100) at room temperature for 20 min with rocking. After per-
(;1.5 mm2) from at least 12 different leaves were cut on a modeling wax
meabilization, samples were washed twice in PBS plus 2% (w/v) BSA and
plate in a drop of 2.5% (w/v) paraformaldehyde and 0.5% (v/v) glutar-
then directly incubated for 30 min at room temperature with EdU Click-IT
aldehyde in 0.06 M Sørensen phosphate buffer, pH 7.2. Samples were
cocktail for detection (Invitrogen) and 20 mg/mL PI (Molecular Probes)
then fixed for 90 min in the same fixing solution. Samples were rinsed in
directly added to the staining solution. The addition of EdU detection
buffer four times for 15 min and dehydrated in increasing concentrations
solution was done according to the manufacturer's instructions. The
of acetone (50, 70, and 90% [v/v]) for 20 min at each step. Subsequently,
samples were then washed twice in PBS, pH 7.4, plus 2% (w/v) BSA,
specimens were gradually infiltrated with increasing concentrations of LR-
transferred to PBS, pH 7.4, and kept at 4°C in the dark for several days
White resin (30, 60, and 100% [w/v]; London Resin) mixed with 90% (v/v)
before imaging. Triton X-100 treatment results in a nuclear stain and is
acetone for a minimum of 3 h per step. Samples were finally embedded in
compatible with EdU costaining. Moreover, it prevents the cell shrinkage
pure, fresh LR-White resin and polymerized at 50°C for 48 h in small
plastic containers under anaerobic conditions. Ultrathin sections (80 nm)
Results) were infiltrated into 4-week-old N. benthamiana leaves as de-
were cut with a Reichert Ultracut S ultramicrotome (Leica Microsystems).
scribed earlier. Expression of the Nb-MEK2 variants was induced with
Immunogold labeling of See1-3xHA was done with ultrathin sections
30 mM DEX 40 to 48 h later (Yang et al., 2001). Treated leaves were collected
on coated nickel grids with the automated immunogold labeling system
;5 h after DEX infiltration. Ground leaf material was thawed in 10 mL of Ex-
Leica EM IGL (Leica Microsystems). The ideal dilutions and incubation
strep buffer (100 mM Tris-HCl, pH 8.0, 5 mM EGTA, 5 mM EDTA, 150 mM
times of the primary monoclonal anti-HA antibody (produced in mouse by
NaCl, 10 mM DTT, 0.5 mM 4-(2-aminoethyl) benzenesulfonyl fluoride hy-
Sigma-Aldrich) and secondary antibodies (goat anti-mouse antibodies
drochloride, 5 mg/mL antipain, 5 mg/mL leupeptin, 50 mM NaF, 1% [v/v]
from British BioCell International) were determined in preliminary studies
Phosphatase Inhibitor Cocktail 1 [Sigma-Aldrich], 0.5% [v/v] Triton X-100,
by evaluating the labeling density after a series of labeling experiments.
and 100 mg/mL avidin) as described previously (Witte et al., 2004). The slurry
The final dilutions of primary and secondary antibodies used in this study
was centrifuged for 10 min at 4°C (15,000g), the supernatant was filtered
showed a minimum background labeling outside the sample with a
through Miracloth, and 0.5 mL of StrepTactin Sepharose (IBA) was added.
maximum specific labeling in the sample. The sections were blocked for
Binding was performed by incubation of this suspension on a rotator for 1 h
20 min with 2% (w/v) BSA (Sigma-Aldrich) in PBS, pH 7.2, and then
at 4°C. The slurry was transferred into a Poly-Prep column (Bio-Rad), and
treated with the primary antibody against See1 3xHA diluted 1:2000 in
the flow-through was discarded. The resin was washed twice with 10 mL of
PBS containing 1% (w/v) BSA. After section washing with PBS containing
W-buffer (100 mM Tris-HCl, pH 8.0, 150 mM NaCl, and 1 mM EDTA). Four
1% (w/v) BSA three times for 5 min, each grid was treated with 10-nm
times 250 mL of E-buffer (100 mM Tris-HCl, pH 8.0, 150 mM NaCl, 1 mM
gold-conjugated secondary antibodies (goat anti-mouse IgG) diluted
EDTA, and 2.5 mM desthiobiotin) was added, and eluates were collected.
1:100 in PBS containing 1% (w/v) BSA for 90 min at room temperature.
The samples were concentrated on a Microcon YM-10 (Millipore) for 30 min
After a short wash in PBS (three times for 5 min) and distilled water (two
at 4°C (13,000g) to a volume of 20 mL and resolved by SDS-PAGE.
times for 5 min), labeled grids were poststained with 2% uranyl-acetateaqueous solution for 15 s and then investigated with a Philips CM10 TEM
Recombinant Protein Preparation
apparatus. Micrographs of randomly photographed immunogold-labeledsections were digitized, and gold particles were counted automatically
The full-length cDNA of maize SGT1 was cloned into the pET-15b vector.
using the software package Cell D with the particle analysis tool
The full-length cDNA of SIPK was cloned into the pGEX-6P-1 vector. The
(Olympus, Life and Material Science Europa) in different visually identified
full-length cDNA of tobacco (Nicotiana tabacum) MEK2DD was cloned into
and manually traced cell structures. The obtained data were statistically
the pGEX-6P-2 vector. Escherichia coli (BL21) cells were induced with 0.5
evaluated using Statistica (Stat-Soft Europe).
mM isopropyl b-D-1-thiogalactopyranoside for Zm-SGT1 at 28°C for 4 hand with 0.25 mM isopropyl b-D-1-thiogalactopyranoside for SIPK and
Coimmunoprecipitation of See1 and SGT1 in N. benthamiana
MEK2DD at 18°C for 4 h. His-tagged recombinant protein was purified usingNi-NTA resin (Qiagen). Glutathione S-transferase-tagged recombinant
proteins were purified using the Glutathione Agarose resin (Sigma-Aldrich).
YFP-HA were heterologously expressed in N. benthamiana. As ex-pression controls, the constructs were separately expressed with the
Targeted Site-Directed Mutagenesis of Maize SGT1
appropriate empty vector. For all experiments, Agrobacterium tumefa-ciens GV3101 was transformed as described previously (Flowers and
The targeted exchange of one or more bases in the plasmids was per-
Vaillancourt, 2005). The transformants were infiltrated into N. ben-
formed by PCR using the QuikChange Multi Site-Directed Mutagenesis kit
thamiana leaves (3 to 4 weeks old) according to Sparkes et al. (2006). Four
(Stratagene). The protocol used for this mutation and base substitution
days after infiltration, the leaves were harvested and ground in liquid
was according to the manufacturer's instructions. Up to three oligonu-
nitrogen. The ground powder was mixed with buffer (50 mM Tris-HCl and
cleotides were designed for the amplification of the entire plasmid, with
150 mM NaCl, pH 7.0). The resulting leaf extract was centrifuged at 3500
one oligonucleotide containing the corresponding mutation(s) in the primer
rpm at 4°C and subsequently sterile-filtered. The protein concentration of
the extract was determined with the Roti-Quant Protein quantificationassay (Carl Roth). To 1 mL of leaf extract containing 2 mg/mL protein, 50m
In Vitro Phosphorylation Assays
L of anti-HA Affinity Matrix (Roche Diagnostics) was added, and samples
were incubated overnight at 4°C on a rotation wheel. The samples were
The purified recombinant 6xHis-Zm-SGT1 or 6xHis-Zm-SGT1AP (5 mg
then centrifuged through Pierce SpinColumns (Thermo Scientific) and
each) was incubated at 30°C for 30 min with SIPK and MEK2DD (2 mg each)
washed once with buffer (50 mM Tris-HCl and 150 mM NaCl, pH 7.0), and
in the reaction buffer (20 mM Tris-HCl, pH 7.5, 40 mM MgCl , and 10 mM
protein was finally eluted by boiling samples in 23 SDS loading buffer for 5
EGTA) or with crude plant protein extracts from the infected maize samples
min. Appropriate amounts of the eluted proteins were separated by SDS-
of seedling leaves and tassel base with U. maydis wild-type SG200,
PAGE, followed by transfer to a nitrocellulose membrane. After electro-
SG200Dsee1, or the overexpression strain with See1 (approximately 20 mg
blotting, the membrane was saturated with 5% (w/v) nonfat dry milk in
each) in the reaction buffer (40 mM Tris-HCl, pH 8.0, 10 mM MgCl , 1 mM
TBS-T (50 mM Tris-HCl, 150 mM NaCl, pH 7.6, and 0.1% [v/v] Tween 20)
DTT, and 0.1% [v/v] Triton X-100). Both buffers contained 50 mM ATP and
for 1 h at room temperature. After blocking, the membrane was washed
1.5 mCi of [g-32P]ATP. The reaction was terminated by adding 33 Laemmli
three times with TBS-T followed by incubation with the primary antibody
sample buffer.
(anti-HA antibody, 1:10,000; anti-c-Myc antibody, 1:5000; Sigma-Aldrich)overnight at 4°C. Membranes were washed three times prior to incubation
Mass Spectrometry Analysis
for 1 h with horseradish peroxidase-conjugated secondary antibody (anti-mouse antibody, 1:5000; Cell Signaling). Signals were detected using
Gel bands containing the proteins of interest were subjected to a standard
SuperSignal West Pico Chemiluminescent Substrate (Thermo Scientific).
proteomic procedure as described in Briefly,reduced and alkylated proteins were subjected to trypsin digestion. Theresulting peptides were eluted from the gel, and phosphopeptide enrich-
In Planta Phosphorylation Assay
ment was performed on SwellGel Gallium-Chelated Discs (Phosphopeptide
The constructs encoding maize SGT1 in the presence or absence of See1
Isolation Kit; Thermo Scientific). Liquid chromatography-mass spectrometry
and also in addition to the earlier mentioned controls (described in
analyses of the peptide mixtures were performed on the Orbitrap Velos
Tumor Induction by Ustilago maydis
spectrometer (Thermo Scientific). The Mascot program was used for
AF, AF494083.1; tobacco SIPK, AF165186; tobacco MEK2, AF325168;
database searches, and tandem mass spectrometry spectra of phos-
maize MAPK kinase 2, NP_001104843.1; maize putative MAPK family
phorylated peptides were also curated manually.
protein, AFW85791.1; maize putative MAPK MPK6, ACG37232.1; maize ABAstimulation MAPK, NP_001152745.1; maize unknown kinase, ACF85409.1.
Microarray Analysis
Supplemental Data
For the microarray experiments, maize plants (cv Early Golden Bantam)
Scoring Scheme for Leaf Tumors Based on
were grown in a phytochamber with the same climatic conditions as
Symptoms Observed at 12 DPI.
described previously. Plants were inoculated in three independent bi-ological replicates with water (mock), SG200, and SG200Dsee1 as de-
SG200Δsee1 Mutant Completes Its Entire
scribed previously in virulence assays for seedling infections (Kämper
et al., 2006; Doehlemann et al., 2008). Infected or mock-inoculated tissue
Formation of Ear Tumors in SG200Δsee1 Mutant
from 15 plants per treatment was harvested at 6 DPI by excising a section
Demonstrates the Independence of Floral Tumors on See1 Activity.
of the third leaf between 1 and 3 cm below the injection holes. For RNAextraction, material from 15 plants per treatment was pooled and ground
Growth of SG200Δsee1 Mutant Is Arrested in
to powder in liquid nitrogen, and RNA was extracted with Trizol reagent
the Mesophyll and Vascular Cell Layers of Leaves.
(Invitrogen). RNA was purified using the RNeasy kit (Qiagen). Agilent
Gene Expression during Maize Colonization
4x44k maize genome arrays were used in three biological replicates, using
with SG200 and SG200Δsee1.
the standard Agilent One Color Microarray-based gene expression
Specificity of the EdU Labeling Assay.
analysis low-input quick Amp labeling protocol. Expression data weresubmitted to the Gene Expression Omnibus
See1 Requirement in Leaf Tumor Formation.
accession number GSE63077). For further experimental details,
Constitutive Overexpression of See1 Results
in Tumors on the Vegetative Parts of Tassels.
Stable Expression of the Ppit2 overex-
Plasmid Constructs and Nucleic Acid Construction
pressed See1-HA Protein in Infected Seedlings and Tassel Base.
For plasmid construction, standard molecular cloning strategies and
Overexpression of See1 Results in Two
techniques were applied (Sambrook et al., 1989). All plasmids generated
Abnormalities in Tassels: Tumors at the Tassel Base and Greenish
and used in this study are listed in Oligonucleotides
that were used for PCR are shown in All restrictionenzymes used in this study were purchased from New England Biolabs. For
See1 Is Transferred to Cells Neighboring to
a detailed description of plasmid constructs, see .
the Transformed Cell.
The isolation of genomic U. maydis DNA was performed as described
Stable Expression of the See1-3xHA Protein
previously (Schulz et al., 1990). All U. maydis strains (
in Infected Tissues.
are derived from SG200 and were generated by insertion of p123
See1 Is Translocated to the Plant Cell
derivatives into the ip locus as described (Loubradou et al., 2001). Isolated
Cytoplasm and Nucleus.
U. maydis transformants were tested by DNA gel blot hybridization toassess single or multiple integration events in the ip locus. For all the
Complementation of Two Yeast sgt1 Cell
cloning work, PCR was performed using Phusion High-Fidelity (New
Cycle Temperature Sensitive Mutants with Maize SGT1.
England Biolabs). The PCR products of the different genes were cleaned
The Deduced SGT1 Protein Sequences of
up before digestion using the Wizard SV Gel and PCR Clean-Up System
Z. mays and N. benthamiana Showing a Multiple Alignment.
(Promega). The vectors were transformed into either DH5a or Top10 cells(Invitrogen) and then plated on YT-agar plates containing a specific se-
The Deduced SIPK Protein Sequences of
lection marker. Plasmids were extracted using the QIAprep system
N. tabacum and a Set of Five Putative Z. mays SIPK Showing
(Qiagen). All plasmid constructs containing amplified gene fragments
a Multiple Alignment.
were sequenced.
Expression Level of the Nt-MEK2, Nb-Zm-SGT1, and Um-See1 Transiently Expressed in N. benthamiana Leaves
Bioinformatics Tools Applied in This Study
to Assess Zm-SGT1 Phosphorylation in Planta.
The Deduced SGT1 Protein Sequences with
Signal peptide prediction was performed with the online program SignalP
Marked Phosphorylation Sites within Zm-SGT1.
4.1 (Protein conserved domainsearch was performed with the online program Pfam
Salicylic Acid Induced Protein Kinase
The mass spectrometry data were analyzed using Mascot
Phosphorylates Zm-SGT1 in Vitro.
Distiller software (version 2.1.1; Matrix Science) and compared with the
Subcellular Localization of the Phosphovar-
NCBInr database using the Mascot database search engine (version 2.1;
iants of Maize SGT1.
Matrix Science), as described in detail in
Complementation of Two Yeast sgt1 CellCycle Temperature-Sensitive Mutants with Maize SGT1 Wild Type,
Accession Numbers
Phosphomimic, and Phosphonull.
Sequence data from this article can be found in the GenBank/EMBL data
In Planta Phosphorylation of Maize SGT1AP.
libraries under the following accession numbers: U. maydis see1,
Differentially Expressed Top GO Terms
XP_758386.1; U. maydis pit2, XP_757522.1; U. maydis ppi, XM_754780.1;
Related to the DNA Synthesis and Cell Differentiation.
maize SGT1.1 of cv Early Golden Bantam, KP789376; maize SGT1.2,ACF84496.1; N. benthamiana SGT1 AY, AY899199.1; N. benthamiana SGT1
Oligonucleotides Used in This Study.
Plasmids Used in This Study.
Brefort, T., Doehlemann, G., Mendoza-Mendoza, A., Reissmann,
S., Djamei, A., and Kahmann, R. (2009). Ustilago maydis as a pathogen.
U. maydis Strains Used in This Study.
Annu. Rev. Phytopathol. 47: 423–445.
Peptides Identified by the Mass Spectrometry.
Brefort, T., Tanaka, S., Neidig, N., Doehlemann, G., Vincon, V., and
Kahmann, R. (2014). Characterization of the largest effector genecluster of Ustilago maydis. PLoS Pathog. 10: e1003866.
Cohen, P. (2000). The regulation of protein function by multisite phos-
Comparison of Differentially Expressed
phorylation—A 25 year update. Trends Biochem. Sci. 25: 596–601.
Genes at 6 dpi In Maize Leaves as Comparison of Mock Infected
Conover, W.J. (1999). Practical Nonparametric Statistics, (New York:
versus the Wild Type-Infected SG200 and Mock-Infected versus
John Wiley & Sons).
SG200Δsee1 Mutant Infected at 6 dpi.
Cui, H., Wang, Y., Xue, L., Chu, J., Yan, C., Fu, J., Chen, M., Innes,
R.W., and Zhou, J.M. (2010). Pseudomonas syringae effector pro-
Comparison of Differentially Expressed
tein AvrB perturbs Arabidopsis hormone signaling by activating
Genes at 6 dpi In Maize Leaves Infected with SG200 and SG200Δsee1
MAP kinase 4. Cell Host Microbe 7: 164–175.
Cuzick, A., Maguire, K., and Hammond-Kosack, K.E. (2009). Lack
Full Mascot Search Results of Zm-SGT1
of the plant signalling component SGT1b enhances disease re-
Mass Spectrometry Analysis.
sistance to Fusarium culmorum in Arabidopsis buds and flowers.
New Phytol. 181: 901–912.
Djamei, A., and Kahmann, R. (2012). Ustilago maydis: Dissecting the
molecular interface between pathogen and plant. PLoS Pathog. 8:
Djamei, A., et al. (2011). Metabolic priming by a secreted fungal ef-
We thank Urs Lahrmann for support with the microarray experiment,
fector. Nature 478: 395–398.
Christoph Hemetsberger and Malgorzata Lichocka for help with confocal
Doehlemann, G., Reissmann, S., Assmann, D., Fleckenstein, M.,
microscopy, and Daniela Assmann for expert technical support. We
and Kahmann, R. (2011). Two linked genes encoding a secreted
thank Nam Hai Chua from The Rockefeller University for the DEX-
effector and a membrane protein are essential for Ustilago maydis-
inducible plasmid pTA7001 and Katsumi Kitagawa for the sgt1 yeast
induced tumour formation. Mol. Microbiol. 81: 751–766.
strains. We thank Michal Dadlez (Institute of Biochemistry and Bio-
Doehlemann, G., Requena, N., Schaefer, P., Brunner, F., O'Connell, R.,
physics) for mass spectrometry analyses. This work was supported by
and Parker, J.E. (2014). Reprogramming of plant cells by filamentous
the Max Planck Society the German Research Founda-
plant-colonizing microbes. New Phytol. 204: 803–814.
tion (Grant DO 1421/3-1), and the Cluster of Excellence on
Doehlemann, G., van der Linde, K., Assmann, D., Schwammbach,
Plant Science. A.R. was supported by a fellowship from the German
D., Hof, A., Mohanty, A., Jackson, D., and Kahmann, R. (2009).
Academic Exchange Service and a Short Term Scientific
Pep1, a secreted effector protein of Ustilago maydis, is required for
Mission grant from the COST FA 1208 program
successful invasion of plant cells. PLoS Pathog. 5: e1000290.
R.H. was supported by the National Science Centre, Poland
Doehlemann, G., Wahl, R., Horst, R.J., Voll, L.M., Usadel, B., Poree,
F., Stitt, M., Pons-Kühnemann, J., Sonnewald, U., Kahmann, R.,and Kämper, J. (2008). Reprogramming a maize plant: Transcrip-tional and metabolic changes induced by the fungal biotroph Usti-
AUTHOR CONTRIBUTIONS
lago maydis. Plant J. 56: 181–195.
A.R., R.H., M.K., and G.D. conceived and designed the experiments.
Dubacq, C., Guerois, R., Courbeyrette, R., Kitagawa, K., and
A.R., L.S., R.H., and B.Z. performed the research. A.R., L.S., R.H., B.Z.,
Mann, C. (2002). Sgt1p contributes to cyclic AMP pathway activ-
M.K., V.W., and G.D. analyzed data. A.R. and G.D. wrote the article. M.K.
ity and physically interacts with the adenylyl cyclase Cyr1p/Cdc35p
and V.W. edited the article.
in budding yeast. Eukaryot. Cell 1: 568–582.
El Oirdi, M., and Bouarab, K. (2007). Plant signalling components
EDS1 and SGT1 enhance disease caused by the necrotrophic
Received August 14, 2014; revised March 3, 2015; accepted March 31,
pathogen Botrytis cinerea. New Phytol. 175: 131–139.
2015; published April 17, 2015.
Flowers, J.L., and Vaillancourt, L.J. (2005). Parameters affecting the
efficiency of Agrobacterium tumefaciens-mediated transformationof Colletotrichum graminicola. Curr. Genet. 48: 380–388.
Gao, L., Kelliher, T., Nguyen, L., and Walbot, V. (2013). Ustilago maydis
reprograms cell proliferation in maize anthers. Plant J. 75: 903–914.
Austin, M.J., Muskett, P., Kahn, K., Feys, B.J., Jones, J.D., and
Garcia-Ranea, J.A., Mirey, G., Camonis, J., and Valencia, A. (2002).
Parker, J.E. (2002). Regulatory role of SGT1 in early R gene-mediated
p23 and HSP20/alpha-crystallin proteins define a conserved se-
plant defenses. Science 295: 2077–2080.
quence domain present in other eukaryotic protein families. FEBS
Azevedo, C., Sadanandom, A., Kitagawa, K., Freialdenhoven, A.,
Lett. 529: 162–167.
Shirasu, K., and Schulze-Lefert, P. (2002). The RAR1 interactor
Gratzner, H.G. (1982). Monoclonal antibody to 5-bromo- and 5-
SGT1, an essential component of R gene-triggered disease resistance.
iododeoxyuridine: A new reagent for detection of DNA replication.
Science 295: 2073–2076.
Science 218: 474–475.
Bao, Z., Yang, H., and Hua, J. (2013). Perturbation of cell cycle
Gray, W.M., Muskett, P.R., Chuang, H.W., and Parker, J.E. (2003).
regulation triggers plant immune response via activation of disease
Arabidopsis SGT1b is required for SCFTIR1-mediated auxin re-
resistance genes. Proc. Natl. Acad. Sci. USA 110: 2407–2412.
sponse. Plant Cell 15: 1310–1319.
Bhavsar, A.P., et al. (2013). The Salmonella type III effector SspH2
Hann, D.R., and Rathjen, J.P. (2007). Early events in the pathoge-
specifically exploits the NLR co-chaperone activity of SGT1 to
nicity of Pseudomonas syringae on Nicotiana benthamiana. Plant J.
subvert immunity. PLoS Pathog. 9: e1003518.
49: 607–618.
Tumor Induction by Ustilago maydis
Hemetsberger, C., Herrberger, C., Zechmann, B., Hillmer, M., and
Noël, L.D., Cagna, G., Stuttmann, J., Wirthmüller, L., Betsuyaku,
Doehlemann, G. (2012). The Ustilago maydis effector Pep1 sup-
S., Witte, C.P., Bhat, R., Pochon, N., Colby, T., and Parker, J.E.
presses plant immunity by inhibition of host peroxidase activity.
(2007). Interaction between SGT1 and cytosolic/nuclear HSC70
PLoS Pathog. 8: e1002684.
chaperones regulates Arabidopsis immune responses. Plant Cell
Heyneke, E., Luschin-Ebengreuth, N., Krajcer, I., Wolkinger, V., Müller,
19: 4061–4076.
M., and Zechmann, B. (2013). Dynamic compartment specific changes
Ökmen, B., and Doehlemann, G. (2014). Inside plant: Biotrophic
in glutathione and ascorbate levels in Arabidopsis plants exposed to
strategies to modulate host immunity and metabolism. Curr. Opin.
different light intensities. BMC Plant Biol. 13: 104.
Plant Biol. 20: 19–25.
Hogenhout, S.A., Van der Hoorn, R.A.L., Terauchi, R., and Kamoun, S.
Peart, J.R., et al. (2002). Ubiquitin ligase-associated protein SGT1 is
(2009). Emerging concepts in effector biology of plant-associated or-
required for host and nonhost disease resistance in plants. Proc.
ganisms. Mol. Plant Microbe Interact. 22: 115–122.
Natl. Acad. Sci. USA 99: 10865–10869.
Horst, R.J., Doehlemann, G., Wahl, R., Hofmann, J., Schmiedl, A.,
Salic, A., and Mitchison, T.J. (2008). A chemical method for fast and
Kahmann, R., Kämper, J., Sonnewald, U., and Voll, L.M. (2010).
sensitive detection of DNA synthesis in vivo. Proc. Natl. Acad. Sci.
Ustilago maydis infection strongly alters organic nitrogen allocation
USA 105: 2415–2420.
in maize and stimulates productivity of systemic source leaves.
Sambrook, J., Fritsch, E.F., and Maniatis, T. (1989). Molecular
Plant Physiol. 152: 293–308.
Cloning: A Laboratory Manual,
(Cold Spring Harbor, NY: Cold
Hoser, R., Zurczak, M., Lichocka, M., Zuzga, S., Dadlez, M., Samuel,
Spring Harbor Laboratory Press).
M.A., Ellis, B.E., Stuttmann, J., Parker, J.E., Hennig, J., and
Schilling, L., Matei, A., Redkar, A., Walbot, V., and Doehlemann, G.
Krzymowska, M. (2013). Nucleocytoplasmic partitioning of tobacco
(2014). Virulence of the maize smut Ustilago maydis is shaped by
N receptor is modulated by SGT1. New Phytol. 200: 158–171.
organ-specific effectors. Mol. Plant Pathol. 15: 780–789.
Jelenska, J., van Hal, J.A., and Greenberg, J.T. (2010). Pseudo-
Schirawski, J., et al. (2010). Pathogenicity determinants in smut fungi
monas syringae hijacks plant stress chaperone machinery for viru-
revealed by genome comparison. Science 330: 1546–1548.
lence. Proc. Natl. Acad. Sci. USA 107: 13177–13182.
Schulz, B., Banuett, F., Dahl, M., Schlesinger, R., Schäfer, W.,
Jones, J.D., and Dangl, J.L. (2006). The plant immune system. Nature
Martin, T., Herskowitz, I., and Kahmann, R. (1990). The b alleles of
444: 323–329.
U. maydis, whose combinations program pathogenic development,
Kämper, J., et al. (2006). Insights from the genome of the biotrophic
code for polypeptides containing a homeodomain-related motif.
fungal plant pathogen Ustilago maydis. Nature 444: 97–101.
Cell 60: 295–306.
Kankanala, P., Czymmek, K., and Valent, B. (2007). Roles for rice
Shirasu, K. (2009). The HSP90-SGT1 chaperone complex for NLR
membrane dynamics and plasmodesmata during biotrophic in-
immune sensors. Annu. Rev. Plant Biol. 60: 139–164.
vasion by the blast fungus. Plant Cell 19: 706–724.
Skibbe, D.S., Doehlemann, G., Fernandes, J., and Walbot, V.
Kelliher, T., and Walbot, V. (2011). Emergence and patterning of the
(2010). Maize tumors caused by Ustilago maydis require organ-
five cell types of the Zea mays anther locule. Dev. Biol. 350: 32–49.
specific genes in host and pathogen. Science 328: 89–92.
Khang, C.H., Berruyer, R., Giraldo, M.C., Kankanala, P., Park, S.Y.,
Song, J., Durrant, W.E., Wang, S., Yan, S., Tan, E.H., and Dong, X.
Czymmek, K., Kang, S., and Valent, B. (2010). Translocation of
(2011). DNA repair proteins are directly involved in regulation of
Magnaporthe oryzae effectors into rice cells and their subsequent
gene expression during plant immune response. Cell Host Microbe
cell-to-cell movement. Plant Cell 22: 1388–1403.
9: 115–124.
Kim, N.H., Kim, D.S., Chung, E.H., and Hwang, B.K. (2014). Pepper
Song, J., Win, J., Tian, M., Schornack, S., Kaschani, F., Ilyas, M.,
suppressor of the G2 allele of skp1 interacts with the receptor-like
van der Hoorn, R.A., and Kamoun, S. (2009). Apoplastic effectors
cytoplasmic kinase1 and type III effector AvrBsT and promotes the
secreted by two unrelated eukaryotic plant pathogens target the
hypersensitive cell death response in a phosphorylation-dependent
tomato defense protease Rcr3. Proc. Natl. Acad. Sci. USA 106:
manner. Plant Physiol. 165: 76–91.
Kitagawa, K., Skowyra, D., Elledge, S.J., Harper, J.W., and Hieter,
Sparkes, I.A., Runions, J., Kearns, A., and Hawes, C. (2006). Rapid,
P. (1999). SGT1 encodes an essential component of the yeast ki-
transient expression of fluorescent fusion proteins in tobacco plants
netochore assembly pathway and a novel subunit of the SCF
and generation of stably transformed plants. Nat. Protoc. 1: 2019–
ubiquitin ligase complex. Mol. Cell 4: 21–33.
Kotogány, E., Dudits, D., Horváth, G.V., and Ayaydin, F. (2010). A rapid
Takahashi, A., Casais, C., Ichimura, K., and Shirasu, K. (2003).
and robust assay for detection of S-phase cell cycle progression in plant
HSP90 interacts with RAR1 and SGT1 and is essential for RPS2-
cells and tissues by using ethynyl deoxyuridine. Plant Methods 6: 5.
mediated disease resistance in Arabidopsis. Proc. Natl. Acad. Sci.
Li, P., et al. (2010). The developmental dynamics of the maize leaf
USA 100: 11777–11782.
transcriptome. Nat. Genet. 42: 1060–1067.
Talarczyk, A., Krzymowska, M., Borucki, W., and Hennig, J. (2002).
Loubradou, G., Brachmann, A., Feldbrügge, M., and Kahmann, R.
Effect of yeast CTA1 gene expression on response of tobacco
(2001). A homologue of the transcriptional repressor Ssn6p antagonizes
plants to tobacco mosaic virus infection. Plant Physiol. 129: 1032–
cAMP signalling in Ustilago maydis. Mol. Microbiol. 40: 719–730.
Mueller, A.N., Ziemann, S., Treitschke, S., Aßmann, D., and
Tanaka, S., Brefort, T., Neidig, N., Djamei, A., Kahnt, J., Vermerris,
Doehlemann, G. (2013). Compatibility in the Ustilago maydis-
W., Koenig, S., Feussner, K., Feussner, I., and Kahmann, R.
maize interaction requires inhibition of host cysteine proteases by
(2014). A secreted Ustilago maydis effector promotes virulence by
the fungal effector Pit2. PLoS Pathog. 9: e1003177.
targeting anthocyanin biosynthesis in maize. eLife 3: e01355.
Mueller, O., Kahmann, R., Aguilar, G., Trejo-Aguilar, B., Wu, A., and
van der Linde, K., Hemetsberger, C., Kastner, C., Kaschani, F., van
de Vries, R.P. (2008). The secretome of the maize pathogen Usti-
der Hoorn, R.A., Kumlehn, J., and Doehlemann, G. (2012). A
lago maydis. Fungal Genet. Biol. 45 (suppl. 1): S63–S70.
maize cystatin suppresses host immunity by inhibiting apoplastic
Mukhtar, M.S., et al. (2011). Independently evolved virulence effec-
cysteine proteases. Plant Cell 24: 1285–1300.
tors converge onto hubs in a plant immune system network. Sci-
Walbot, V., and Skibbe, D.S. (2010). Maize host requirements for
ence 333: 596–601.
Ustilago maydis tumor induction. Sex. Plant Reprod. 23: 1–13.
Wang, K., Uppalapati, S.R., Zhu, X., Dinesh-Kumar, S.P., and
damage responses to potentiate plant immunity. Mol. Cell 52: 602–
Mysore, K.S. (2010). SGT1 positively regulates the process of
plant cell death during both compatible and incompatible plant-
Yang, K.Y., Liu, Y., and Zhang, S. (2001). Activation of a mitogen-
pathogen interactions. Mol. Plant Pathol. 11: 597–611.
activated protein kinase pathway is involved in disease resistance in
Wen, W., Meinkoth, J.L., Tsien, R.Y., and Taylor, S.S. (1995).
tobacco. Proc. Natl. Acad. Sci. USA 98: 741–746.
Identification of a signal for rapid export of proteins from the nu-
Zhang, M., Kadota, Y., Prodromou, C., Shirasu, K., and Pearl, L.H.
cleus. Cell 82: 463–473.
(2010). Structural basis for assembly of Hsp90-Sgt1-CHORD pro-
Witte, C.P., Noël, L.D., Gielbert, J., Parker, J.E., and Romeis, T.
tein complexes: Implications for chaperoning of NLR innate immu-
(2004). Rapid one-step protein purification from plant material using
nity receptors. Mol. Cell 39: 269–281.
the eight-amino acid StrepII epitope. Plant Mol. Biol. 55: 135–147.
Zhang, S., and Liu, Y. (2001). Activation of salicylic acid-induced
Yan, S., Wang, W., Marqués, J., Mohan, R., Saleh, A., Durrant,
protein kinase, a mitogen-activated protein kinase, induces multiple
W.E., Song, J., and Dong, X. (2013). Salicylic acid activates DNA
defense responses in tobacco. Plant Cell 13: 1877–1889.
Source: http://plantpath.ibb.waw.pl/pdf/ustilago.pdf
See discussions, stats, and author profiles for this publication at: Article · January 2016DOI: 10.1016/j.foodres.2016.01.012 6 authors, including: 25 PUBLICATIONS 296 CITATIONS 99 PUBLICATIONS 1,052 CITATIONS 325 PUBLICATIONS 5,230 CITATIONS 318 PUBLICATIONS 4,389 CITATIONS
The Copyright in this manual vests exclusively with The Titan Trust. All rights are reserved.No part of the manual may be modified, added to, reproduced, transmitted, transcribed,stored in a retrievable system or translated into any language in any form or by any means,electronic, mechanical, magnetic, optical, chemical, manual, or otherwise, in whole or inpart, without the prior consent of The Titan Trust. Any unauthorised use of this manual, inwhole or in part, will result in legal action being taken against such user.








